Common multiple comparison procedures illustrated using graphicalMCP
Source:vignettes/graph-examples.Rmd
graph-examples.RmdIntroduction
In confirmatory clinical trials, regulatory guidelines mandate the strong control of the family-wise error rate at a prespecified level . Many multiple comparison procedures (MCPs) have been proposed for this purpose. The graphical approaches are a general framework that include many common MCPs as special cases. In this vignette, we illustrate how to use graphicalMCP to perform some common MCPs.
To test hypotheses using a graphical MCP, each hypothesis receives a weight (called hypothesis weight), where . From to , there could be a directed and weighted edge , which means that when is rejected, its hypothesis weight will be propagated (or transferred) to and determines how much of the propagation. We also require and .
Bonferroni-based procedures
Weighted Bonferroni test
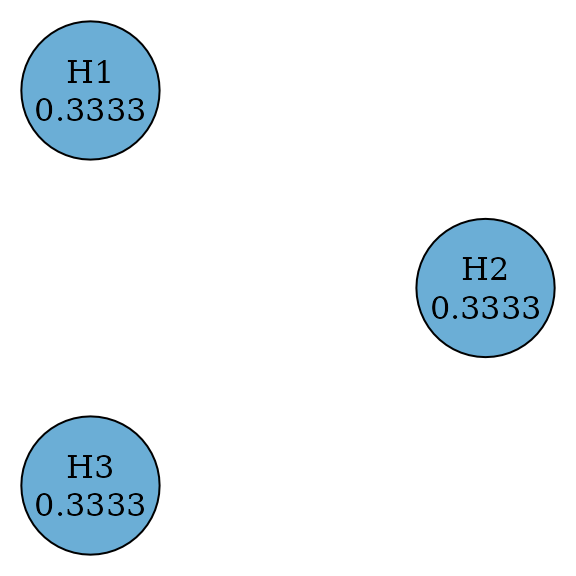
A weighted Bonferroni test splits among hypotheses by testing every hypothesis at a significance level of . Thus it rejects a hypothesis if its p-value is less than or equal to its significance level. When for all , this means an equal split and the test is the Bonferroni test. There is no propagation between any pair of hypothesis.
set.seed(1234)
alpha <- 0.025
m <- 3
bonferroni_graph <- bonferroni(rep(1 / m, m))
# transitions <- matrix(0, m, m)
# bonferroni_graph <- graph_create(rep(1 / m, m), transitions)
plot(
bonferroni_graph,
layout = igraph::layout_in_circle(
as_igraph(bonferroni_graph),
order = c(2, 1, 3)
),
vertex.size = 70
)
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
bonferroni_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3
#> TRUE FALSE FALSEHolm Procedure
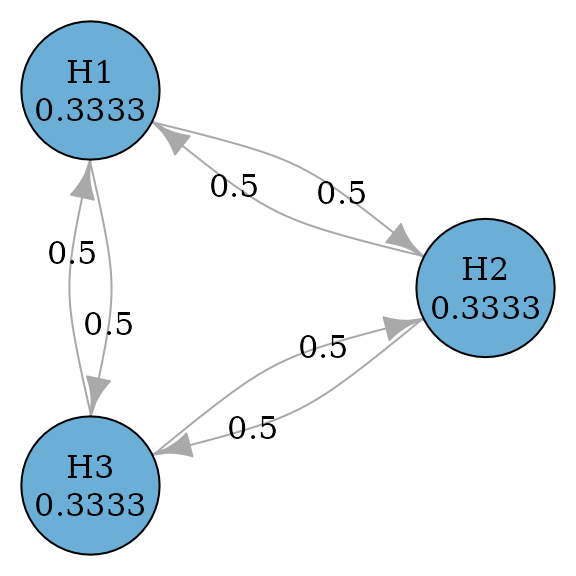
Holm (or Bonferroni-Holm) procedures improve over Bonferroni tests by allowing propagation (Holm 1979). In other words, transition weights between hypotheses may not be zero. So it is uniformly more powerful than Bonferroni tests.
set.seed(1234)
alpha <- 0.025
m <- 3
holm_graph <- bonferroni_holm(rep(1 / m, m))
# transitions <- matrix(1 / (m - 1), m, m)
# diag(transitions) <- 0
# holm_graph <- graph_create(rep(1 / m, m), transitions)
plot(holm_graph,
layout = igraph::layout_in_circle(
as_igraph(holm_graph),
order = c(2, 1, 3)
),
vertex.size = 70
)
p_values <- runif(m, 0, alpha)
test_results <- graph_test_shortcut(holm_graph, p = p_values, alpha = alpha)
test_results$outputs$rejected
#> H1 H2 H3
#> TRUE FALSE FALSEFixed sequence procedure
Fixed sequence (or hierarchical) procedures pre-specify an order of testing [Maurer, Hothorn, and Lehmacher (1995);westfall-2001-optimally]. For example, the procedure will test first. If it is rejected, it will test ; otherwise the testing stops. If is rejected, it will test ; otherwise the testing stops. For each hypothesis, it will be tested at the full level, when it can be tested.
set.seed(1234)
alpha <- 0.025
m <- 3
fixed_sequence_graph <- fixed_sequence(m)
# transitions <- rbind(
# c(0, 1, 0),
# c(0, 0, 1),
# c(0, 0, 0)
# )
# fixed_sequence_graph <- graph_create(c(1, 0, 0), transitions)
plot(fixed_sequence_graph, nrow = 1, asp = 0.05, vertex.size = 40)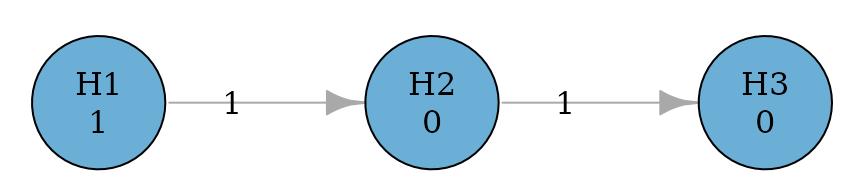
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
fixed_sequence_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3
#> TRUE TRUE TRUEFallback procedure
Fallback procedures have one-way propagation (like fixed sequence procedures) but allow hypotheses to be tested at different significance levels (Wiens 2003).
set.seed(1234)
alpha <- 0.025
m <- 3
fallback_graph <- fallback(rep(1 / 3, 3))
# transitions <- rbind(
# c(0, 1, 0),
# c(0, 0, 1),
# c(0, 0, 0)
# )
# fallback_graph <- graph_create(rep(1 / 3, 3), transitions)
plot(fallback_graph, nrow = 1, asp = 0.05, vertex.size = 40)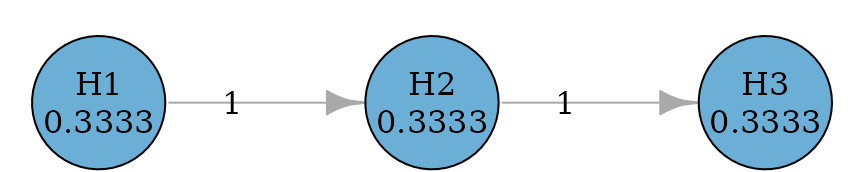
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
fallback_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3
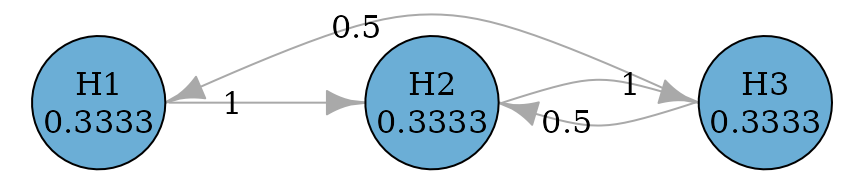
#> TRUE TRUE TRUEFurther they can be improved to allow propagation from later
hypotheses to earlier hypotheses, because it is possible that a later
hypothesis is rejected before an earlier hypothesis can be rejected.
There are two versions of improvement: fallback_improved_1
due to (Wiens and Dmitrienko 2005) and
fallback_improved_2 due to (Bretz et
al. 2009) respectively.
set.seed(1234)
alpha <- 0.025
m <- 3
fallback_improved_1_graph <- fallback_improved_1(rep(1 / 3, 3))
# hypotheses <- rep(1 / 3, 3)
# transitions <- rbind(
# c(0, 1, 0),
# c(0, 0, 1),
# c(hypotheses[seq_len(m - 1)] / sum(hypotheses[seq_len(m - 1)]), 0)
# )
# fallback_improved_1_graph <- graph_create(rep(1 / 3, 3), transitions)
plot(
fallback_improved_1_graph,
nrow = 1,
asp = 0.05,
vertex.size = 40,
edge_curves = c("pairs" = 7, "H3|H1" = -10)
)
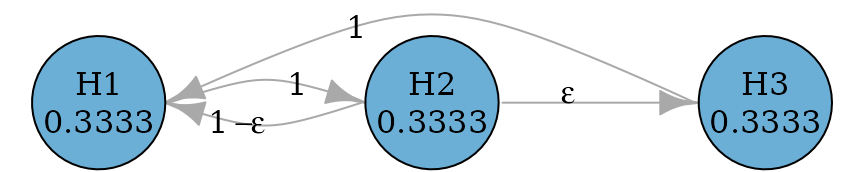
epsilon <- 0.0001
fallback_improved_2_graph <- fallback_improved_2(rep(1 / 3, 3), epsilon)
# hypotheses <- rep(1 / 3, 3)
# transitions <- rbind(
# c(0, 1, 0),
# c(1 - epsilon, 0, epsilon),
# c(1, 0, 0)
# )
# fallback_improved_2_graph <- graph_create(rep(1 / 3, 3), transitions)
plot(
fallback_improved_2_graph,
nrow = 1,
asp = 0.05,
eps = 0.0001,
edge_curves = c("pairs" = 7, "H3|H1" = -10),
vertex.size = 40
)
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
fallback_improved_2_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3
#> TRUE TRUE TRUESerial gatekeeping procedure
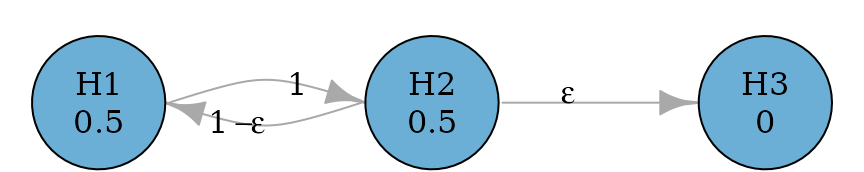
Serial gatekeeping procedures involve ordered multiple families of hypotheses, where all hypotheses of a family of hypotheses must be rejected before proceeding in the test sequence. The example below considers a primary family consisting of two hypotheses and and a secondary family consisting of a single hypothesis . In the primary family, the Holm procedure is applied. If both and are rejected, can be tested at level ; otherwise cannot be rejected. To allow the conditional propagation to , an edge is used from to . It has a very small transition weight so that propagates most of its hypothesis weight to (if not already rejected) and retains a small (non-zero) weight for so that if has been rejected, all hypothesis weight of will be propagated to . Here is assigned to be 0.0001 and in practice, the value could be adjusted but it should be much smaller than the smallest p-value observed.
set.seed(1234)
alpha <- 0.025
m <- 3
epsilon <- 0.0001
transitions <- rbind(
c(0, 1, 0),
c(1 - epsilon, 0, epsilon),
c(0, 0, 0)
)
serial_gatekeeping_graph <- graph_create(c(0.5, 0.5, 0), transitions)
plot(
serial_gatekeeping_graph,
nrow = 1,
asp = 0.05,
eps = 0.0001,
edge_curves = c("pairs" = 7, "H3|H1" = -10),
vertex.size = 40
)
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
serial_gatekeeping_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3
#> TRUE TRUE TRUEParallel gatekeeping procedure
Parallel gatekeeping procedures also involve multiple ordered families of hypotheses, where any null hypotheses of a family of hypotheses must be rejected before proceeding in the test sequence (Dmitrienko, Offen, and Westfall 2003). The example below considers a primary family consisting of two hypotheses and and a secondary family consisting of two hypotheses and . In the primary family, the Bonferroni test is applied. If any of and is rejected, and can be tested at level using the Holm procedure; if both and are rejected, and can be tested at level using the Holm procedure; otherwise and cannot be rejected.
set.seed(1234)
alpha <- 0.025
m <- 4
transitions <- rbind(
c(0, 0, 0.5, 0.5),
c(0, 0, 0.5, 0.5),
c(0, 0, 0, 1),
c(0, 0, 1, 0)
)
parallel_gatekeeping_graph <- graph_create(c(0.5, 0.5, 0, 0), transitions)
plot(parallel_gatekeeping_graph, vertex.size = 70)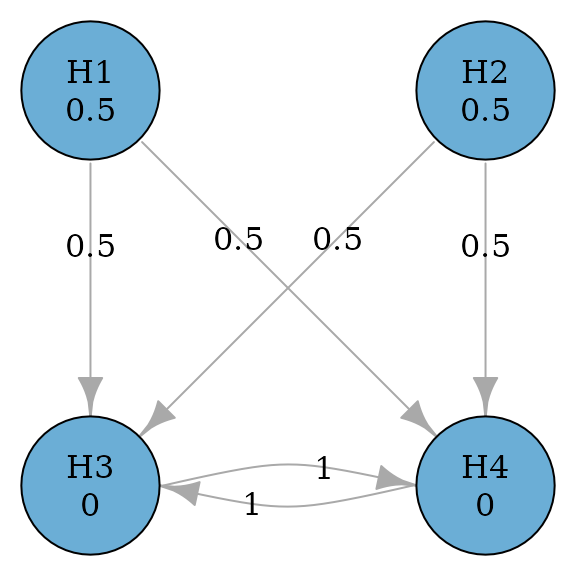
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
parallel_gatekeeping_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3 H4
#> TRUE FALSE FALSE FALSEThe above parallel gatekeeping procedure can be improved by adding edges from secondary hypotheses to primary hypotheses, because it is possible that both secondary hypotheses are rejected but there is still a remaining primary hypothesis not rejected (Bretz et al. 2009).
set.seed(1234)
alpha <- 0.025
m <- 4
epsilon <- 0.0001
transitions <- rbind(
c(0, 0, 0.5, 0.5),
c(0, 0, 0.5, 0.5),
c(epsilon, 0, 0, 1 - epsilon),
c(0, epsilon, 1 - epsilon, 0)
)
parallel_gatekeeping_improved_graph <-
graph_create(c(0.5, 0.5, 0, 0), transitions)
plot(parallel_gatekeeping_improved_graph, eps = 0.0001, vertex.size = 70)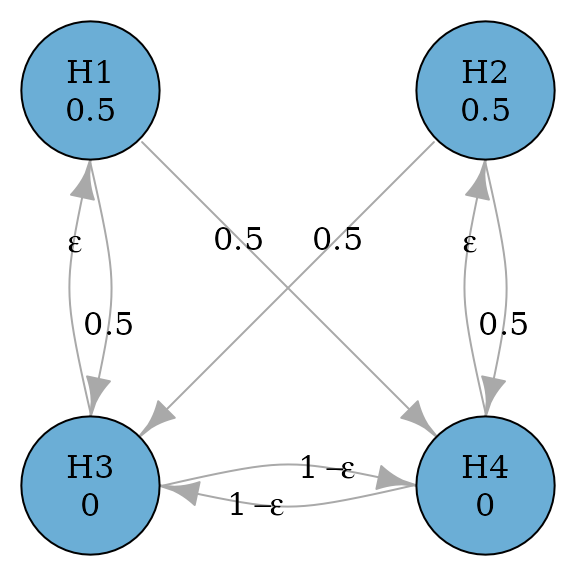
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
parallel_gatekeeping_improved_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3 H4
#> TRUE FALSE FALSE FALSESuccessive procedure
Successive procedures incorporate successive relationships between hypotheses. For example, the secondary hypothesis is not tested until the primary hypothesis has been rejected. This is similar to using the fixed sequence procedure as a component of a graph. The example below considers two primary hypotheses and and two secondary hypotheses and . Primary hypotheses and receive the equal hypothesis weight of 0.5; secondary hypotheses and receive the hypothesis weight of 0. A secondary hypothesis can be tested only if the corresponding primary hypothesis has been rejected. This represents the successive relationships between and , and and , respectively (Maurer, Glimm, and Bretz 2011). If both and are rejected, their hypothesis weights are propagated to and , and vice versa.
set.seed(1234)
alpha <- 0.025
m <- 4
simple_successive_graph <- simple_successive_1()
# transitions <- rbind(
# c(0, 0, 1, 0),
# c(0, 0, 0, 1),
# c(0, 1, 0, 0),
# c(1, 0, 0, 0)
# )
# simple_successive_graph <- graph_create(c(0.5, 0.5, 0, 0), transitions)
plot(simple_successive_graph, layout = "grid", nrow = 2, vertex.size = 70)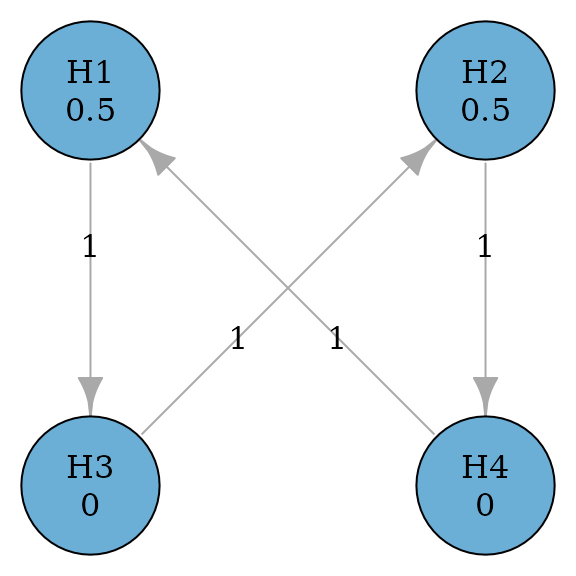
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
simple_successive_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3 H4
#> TRUE FALSE FALSE FALSEThe above graph could be generalized to allow propagation between primary hypotheses (Maurer, Glimm, and Bretz 2011). A general successive graph is illustrate below with a variable to determine the propagation between and .
set.seed(1234)
alpha <- 0.025
m <- 4
successive_var <- simple_successive_var <- function(gamma) {
graph_create(
c(0.5, 0.5, 0, 0),
rbind(
c(0, gamma, 1 - gamma, 0),
c(gamma, 0, 0, 1 - gamma),
c(0, 1, 0, 0),
c(1, 0, 0, 0)
)
)
}
successive_var_graph <- successive_var(0.5)
plot(successive_var_graph, layout = "grid", nrow = 2, vertex.size = 70)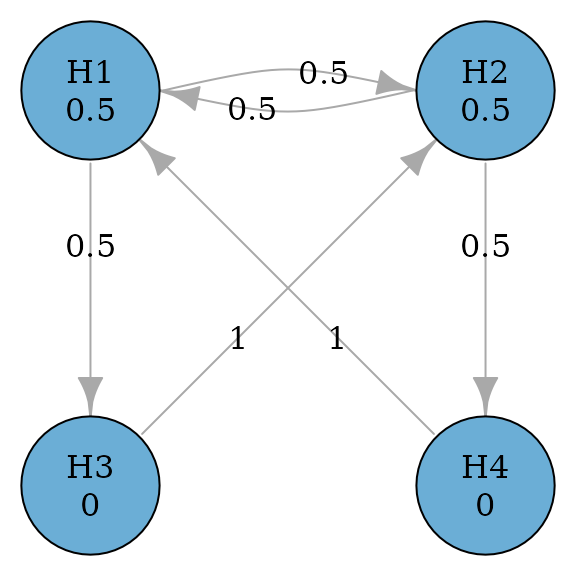
p_values <- runif(m, 0, alpha)
test_results <-
graph_test_shortcut(
successive_var_graph,
p = p_values,
alpha = alpha
)
test_results$outputs$rejected
#> H1 H2 H3 H4
#> TRUE TRUE FALSE FALSESimes-based procedures
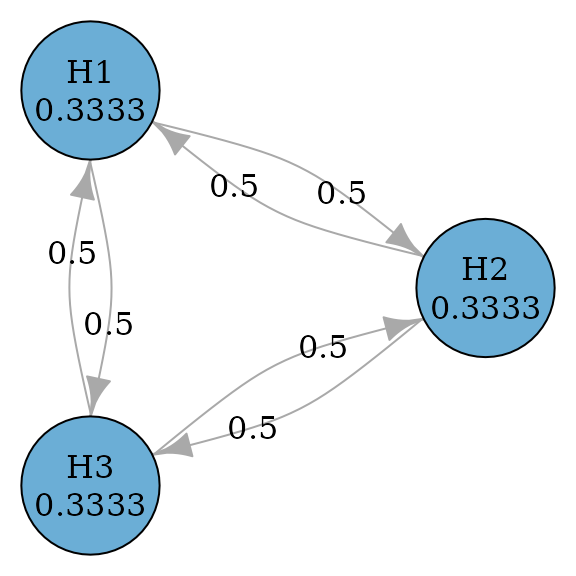
Hommel procedure
Hommel procedure (Hommel 1988) is a
closed test procedure which uses Simes tests for every intersection
hypothesis. According to Xi and Bretz
(2019), the graph for Hommel procedures is the same as the graph
for Holm procedures. Thus to perform Hommel procedure, we just need to
specify test_type to be simes.
set.seed(1234)
alpha <- 0.025
m <- 3
hommel_graph <- bonferroni_holm(rep(1 / m, m))
plot(
hommel_graph,
layout = igraph::layout_in_circle(
as_igraph(hommel_graph),
order = c(2, 1, 3)
),
vertex.size = 70
)
p_values <- runif(m, 0, alpha)
test_results <- graph_test_closure(
hommel_graph,
p = p_values,
alpha = alpha,
test_types = "simes"
)
test_results$outputs$rejected
#> H1 H2 H3
#> TRUE TRUE TRUEParametric procedures
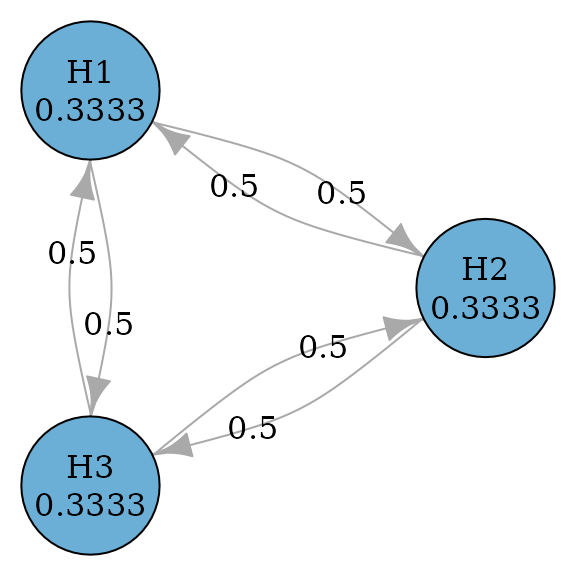
Dunnett procedure
Step-down Dunnett procedures are a closed test procedure and an
improvement from Holm procedures by incorporating the correlation
structure between test statistics (Dunnett and
Tamhane 1991). Thus they are the same graph as Holm procedures.
Assume an equi-correlated case, where the correlation between any pair
of test statistics is the same, e.g., 0.5. Then we can perform the
step-down Dunnett procedure by specifying test_type to be
parametric and providing the correlation matrix.
set.seed(1234)
alpha <- 0.025
m <- 3
dunnett_graph <- bonferroni_holm(rep(1 / m, m))
plot(
dunnett_graph,
layout = igraph::layout_in_circle(
as_igraph(dunnett_graph),
order = c(2, 1, 3)
),
vertex.size = 70
)
p_values <- runif(m, 0, alpha)
corr <- matrix(0.5, m, m)
diag(corr) <- 1
test_results <- graph_test_closure(
dunnett_graph,
p = p_values, alpha = alpha,
test_types = "parametric",
test_corr = list(corr)
)
test_results$outputs$rejected
#> H1 H2 H3
#> TRUE FALSE FALSE