{workr}: a very simple R data pipeline
Phuse Connect 2026
2026-03-26
Agenda
- What is {workr}?
- Why {workr}?
- {workr} ❤️ {pharmaverse}
- {workr} Apps
What is {workr}?
A very simple R data pipeline
{workr} workflows are YAML files easily parsed to an R list
list meta captures metadata
steps are function calls
meta can be used in steps via params
workr::RunWorkflow()
Inputs
- workflow (YAML file parsed to a
list) - initial data (
lData)- available to all
stepsand can be updated by each step
- available to all
Output
- Returns result of last step by default
- If
bReturnResult = FALSEreturns full workflow object with all intermediate outputs
workr::demoApp()
workr::demoApp()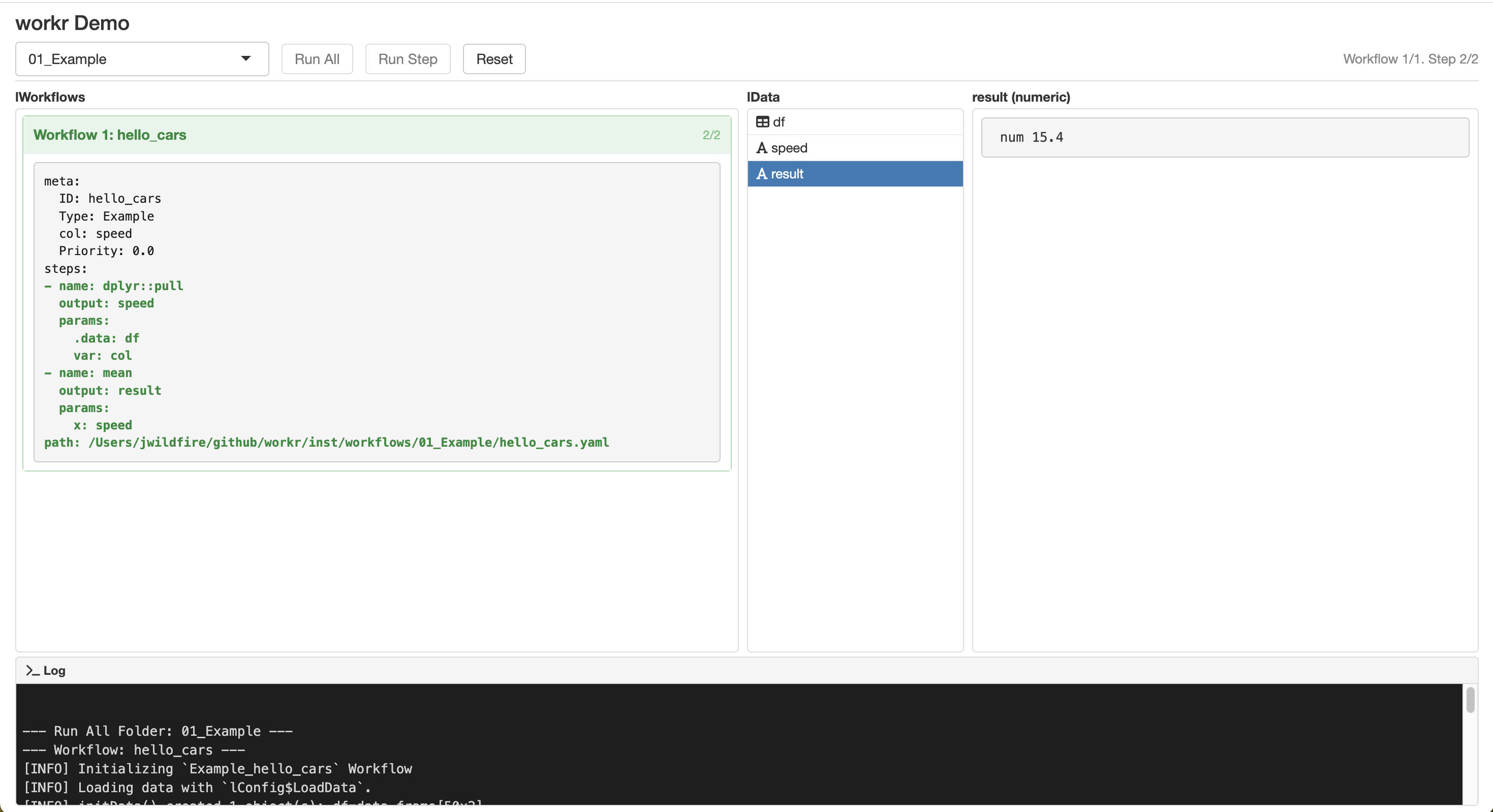
workr::RunWorkflows()
- Convenience function to run multiple workflows in sequence
- Output of each workflow added to
lDatafor the next workflow
workr::demoApp()
workr::demoApp()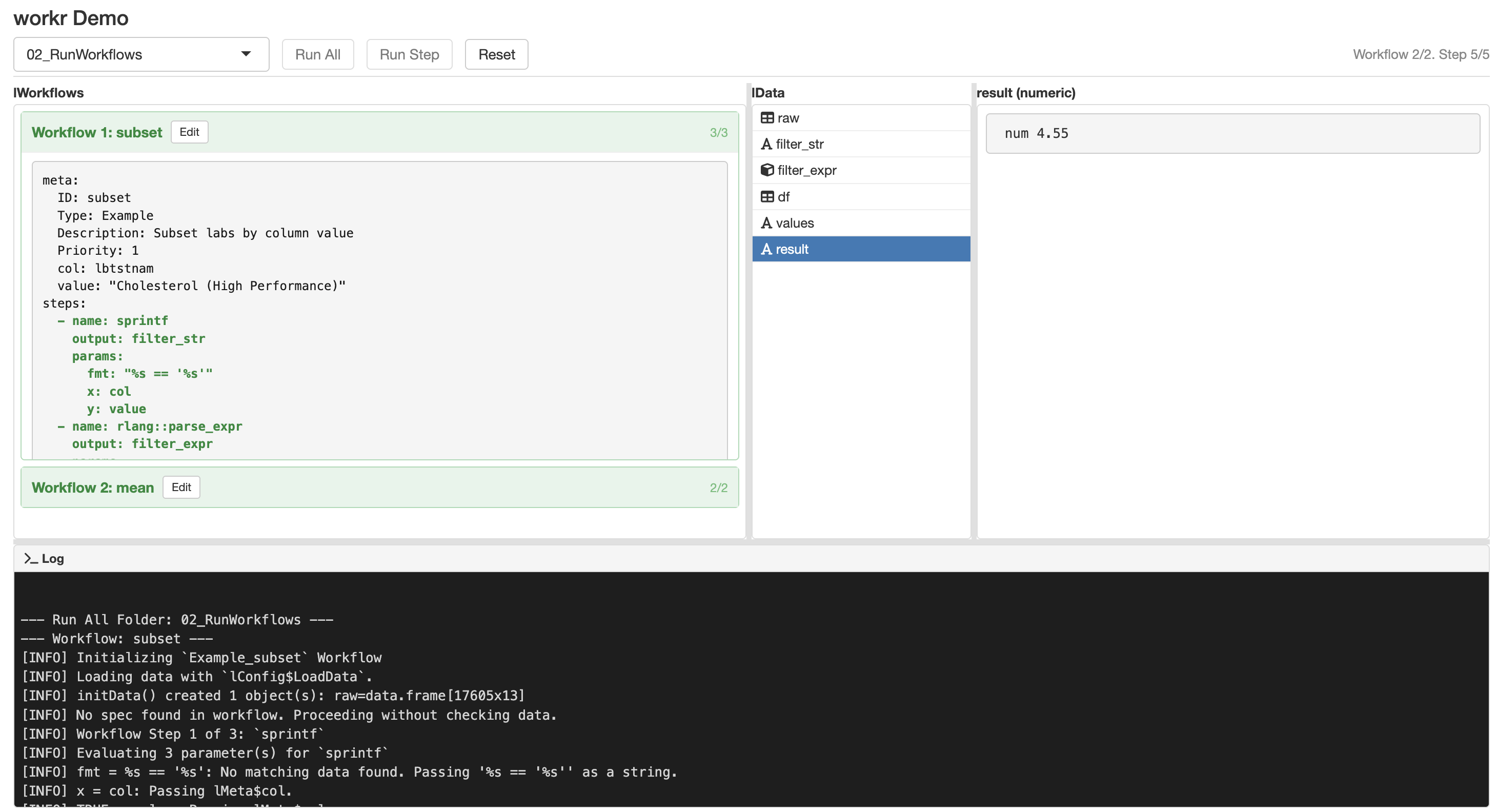
That’s really about it
- Minimal mental model
- Easy to read / debug
- Surprisingly scalable
Why {workr}?
Scalable Clinical Trial Operations
From Safety Monitoring …
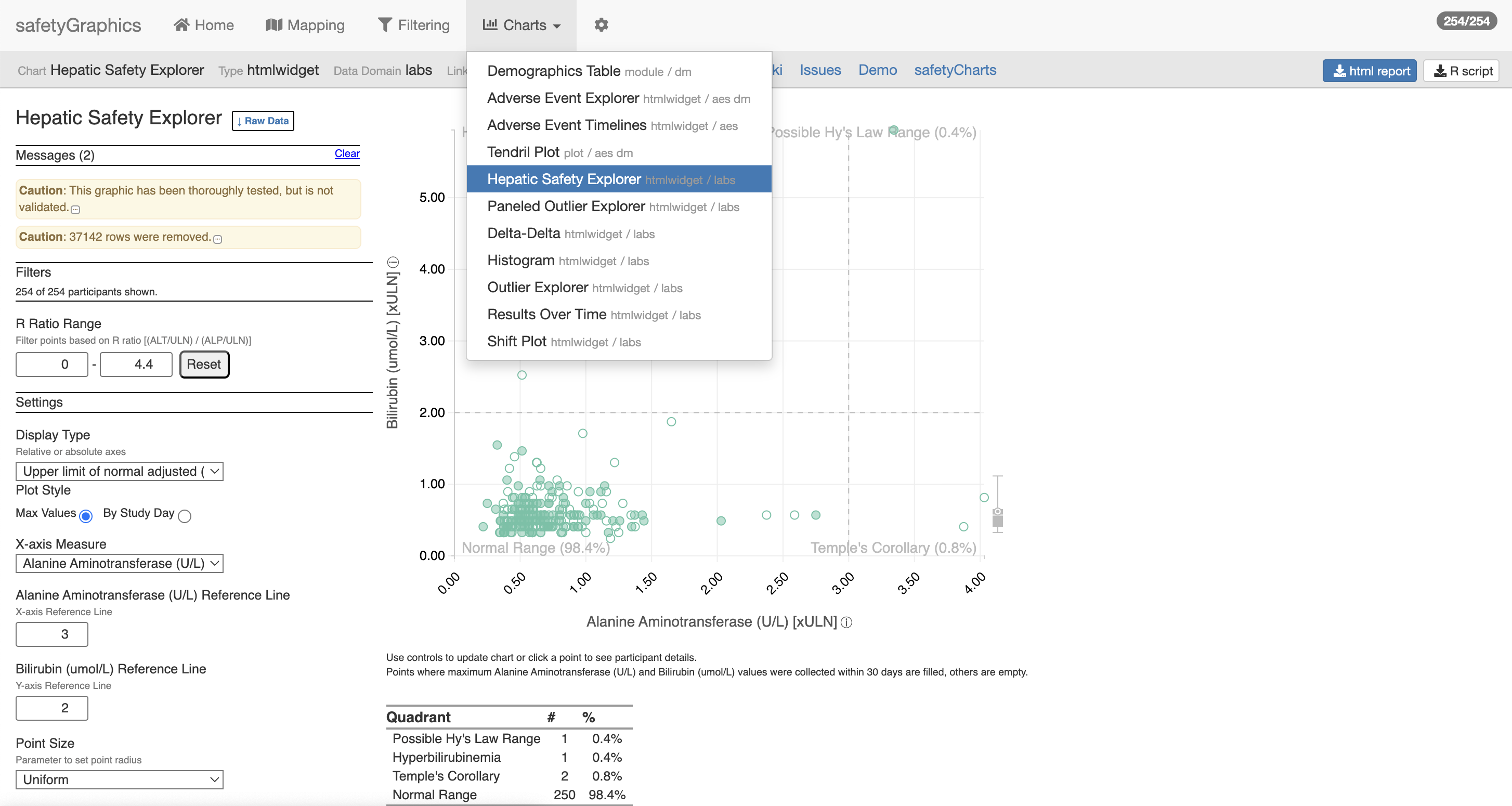
With gnarly data pipelines …

To Risk-Based Quality Monitoring
{gsm}(Good Statistical Monitoring) is a qualified set of R packages that provides a GxP framework for central monitoring in clinical trials.- Designed to support a variety of business processes, from manually executed RBQM reporting to enterprise-wide monitoring platforms.
Repeatability and scalability are key
- Data pipeline: 30 studies × monthly snapshots × 15 metrics × 5 steps
- ~27,000 metrics / year
But Science is Hard
Clinical trials are complex
- Study designs vary
- Metrics need tweaks
- Study-level (and sometimes metric-level) customization required
🚫 Custom Study Scripts 🚫
- Needed reusable pipelines / workflows
- Existing tools felt a bit complicated (
targets,glue, etc.) - So we created
gsm::RunWorkflow()in April 2022 - Migrated to
{workr}for better modularity and extensibility
{gsm} metrics
{gsm} provides a framework that allows users to assess and visualize site-level risk in clinical trial data.
12 Core Site KRIs
- Adverse Event Reporting Rate
- Serious Adverse Event Reporting Rate
- Non-important Protocol Deviation Rate
- Important Protocol Deviation Rate
- Grade 3+ Lab Abnormality Rate
- Study Discontinuation Rate
- Treatment Discontinuation Rate
- Query Rate
- Outstanding Query Rate
- Outstanding Data Entry Rate
- Data Change Rate
- Screen Failure Rate
Highly Standardized Metric Workflows
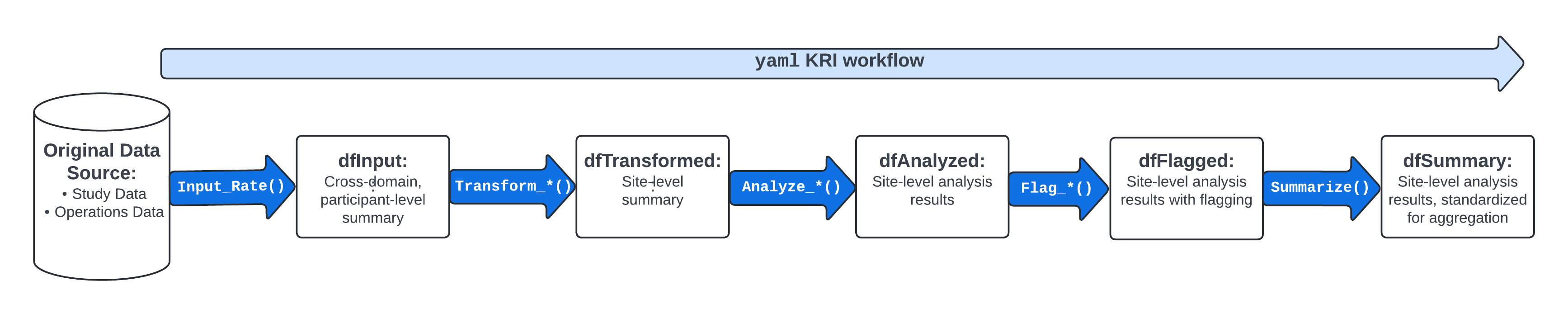
workr::demoApp()
workr::demoApp()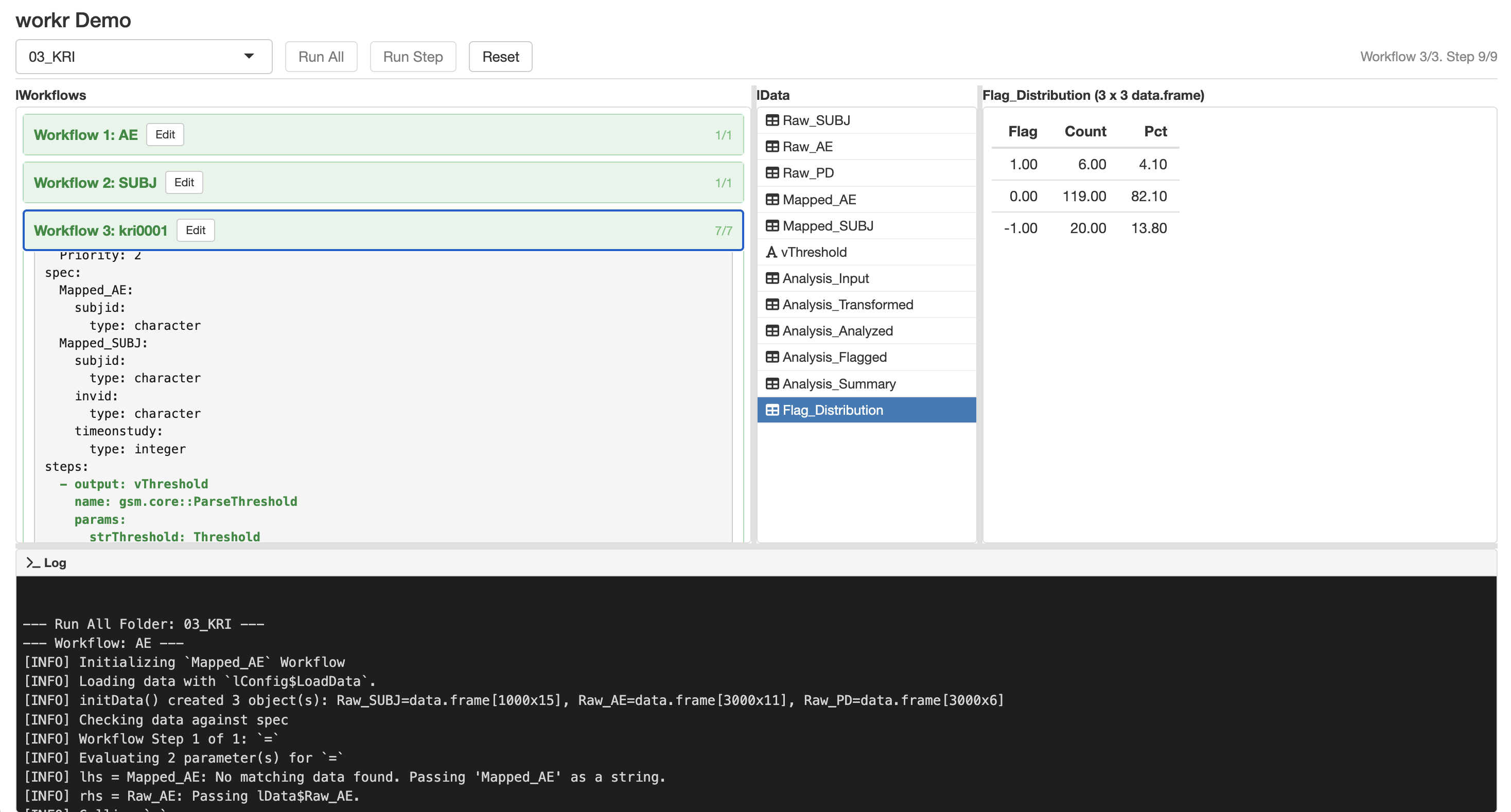
{gsm} extensions Reporting and Visualization Layer
Site KRI Report
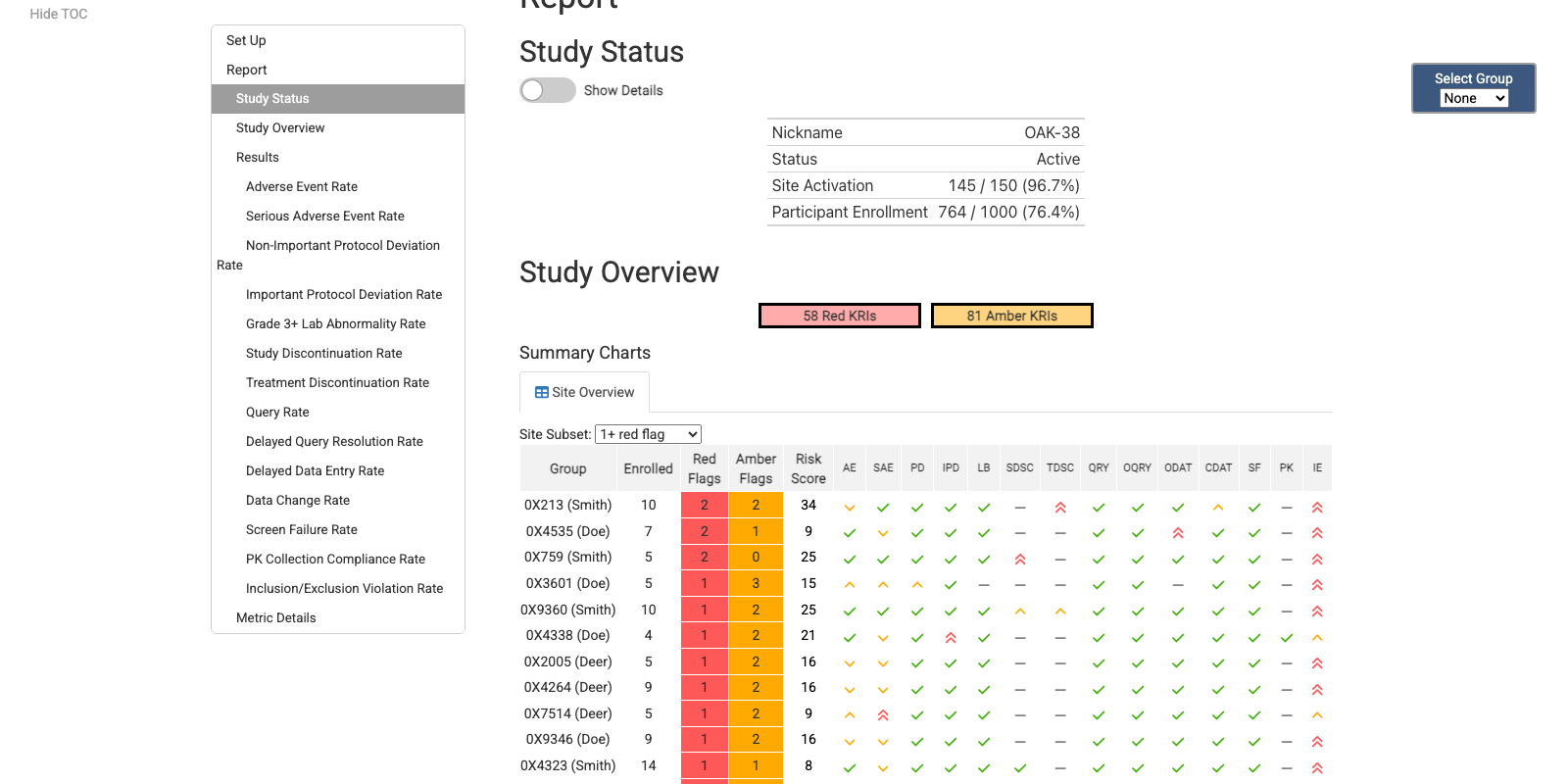
Country KRI Report
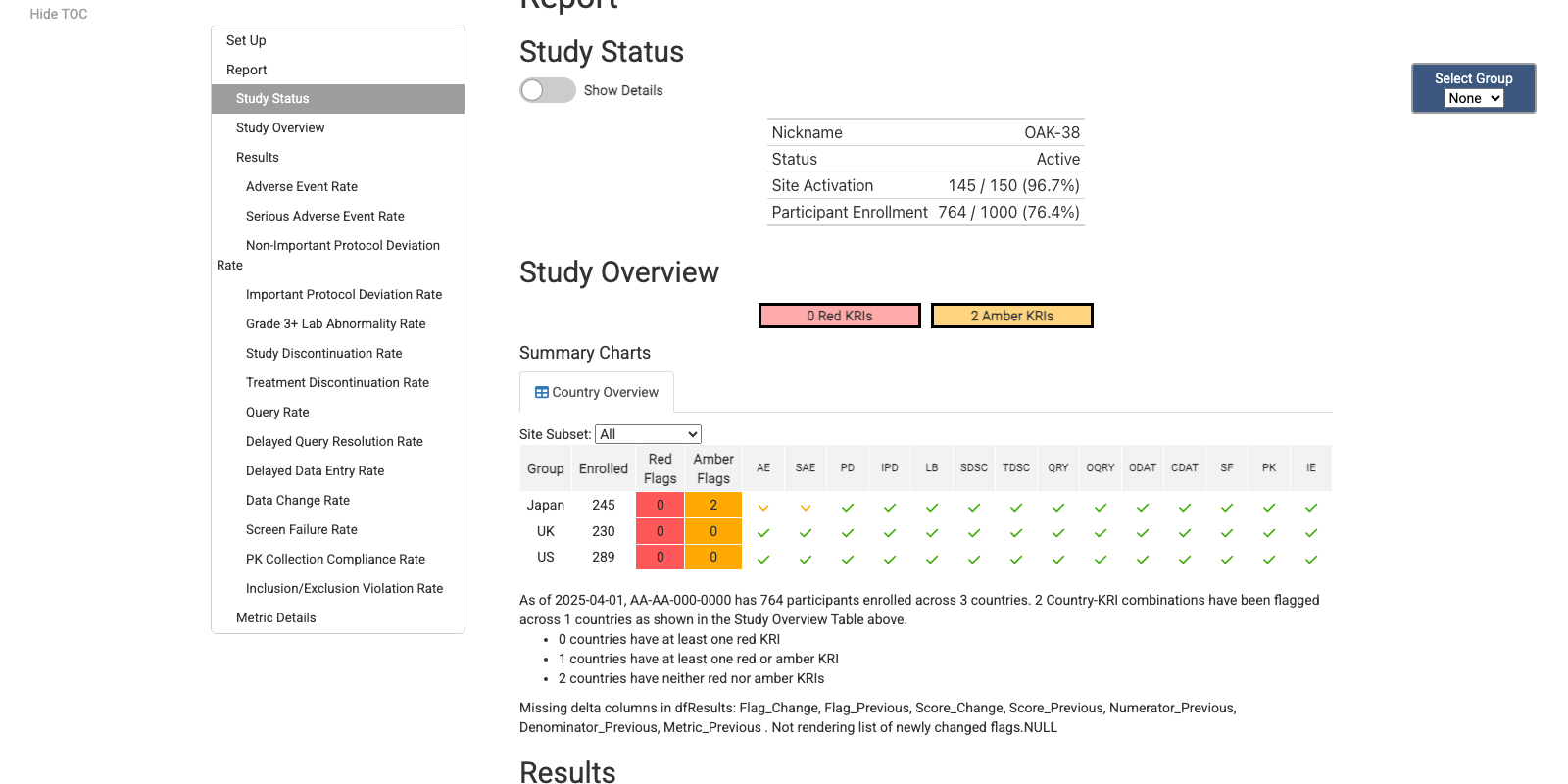
Cross-Study KRI Report
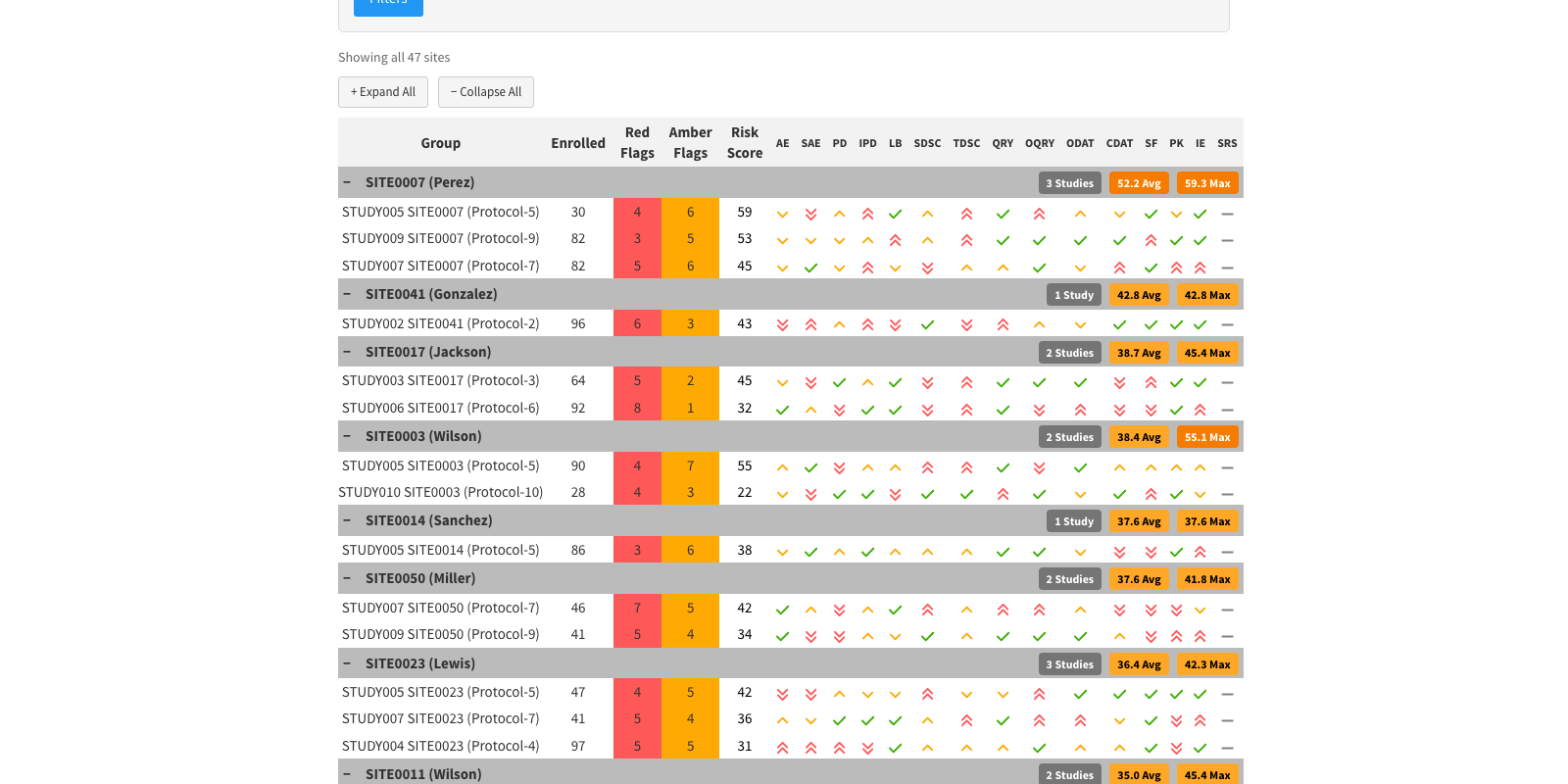
Eligibility Report
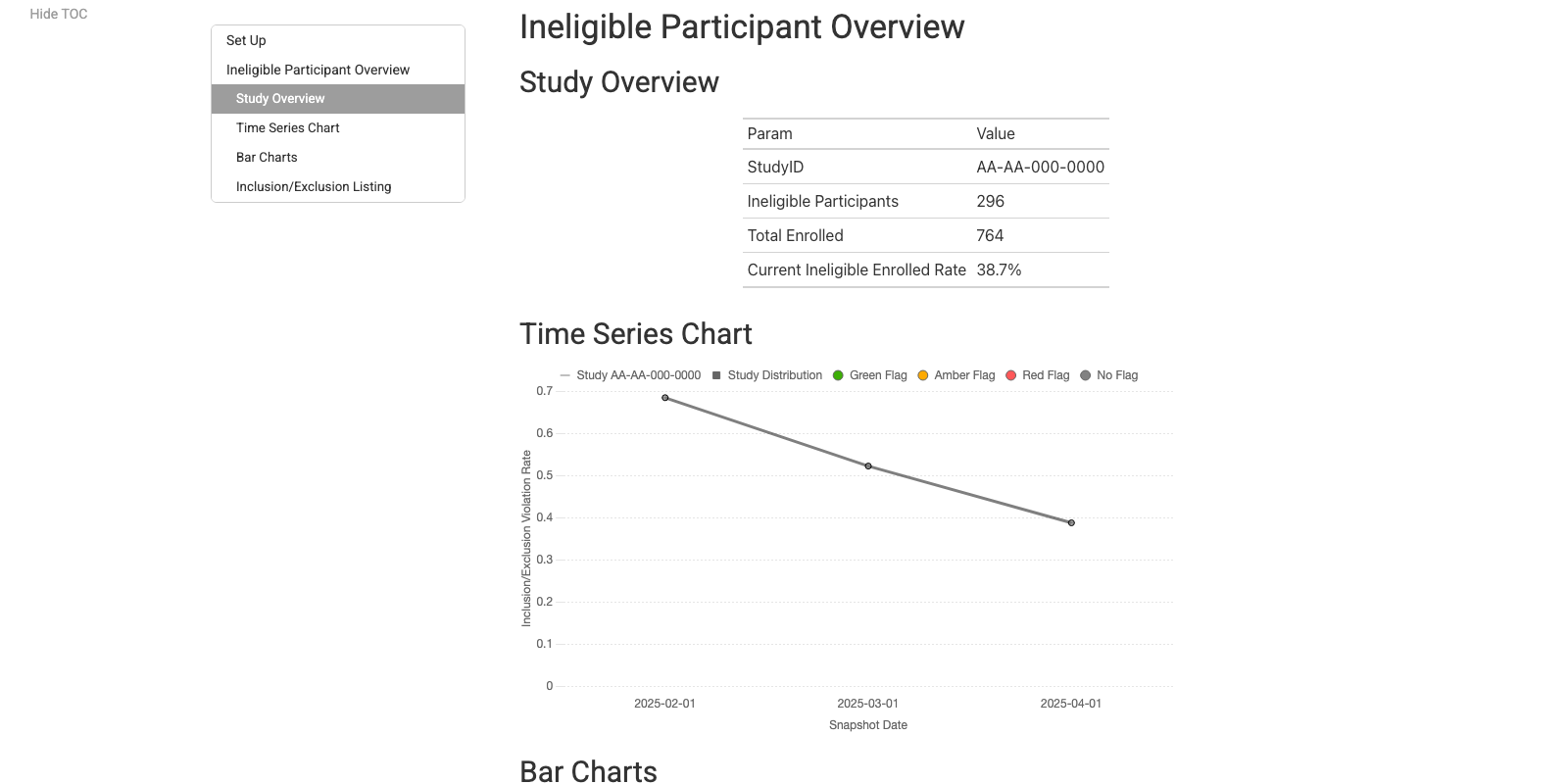
QTL Report
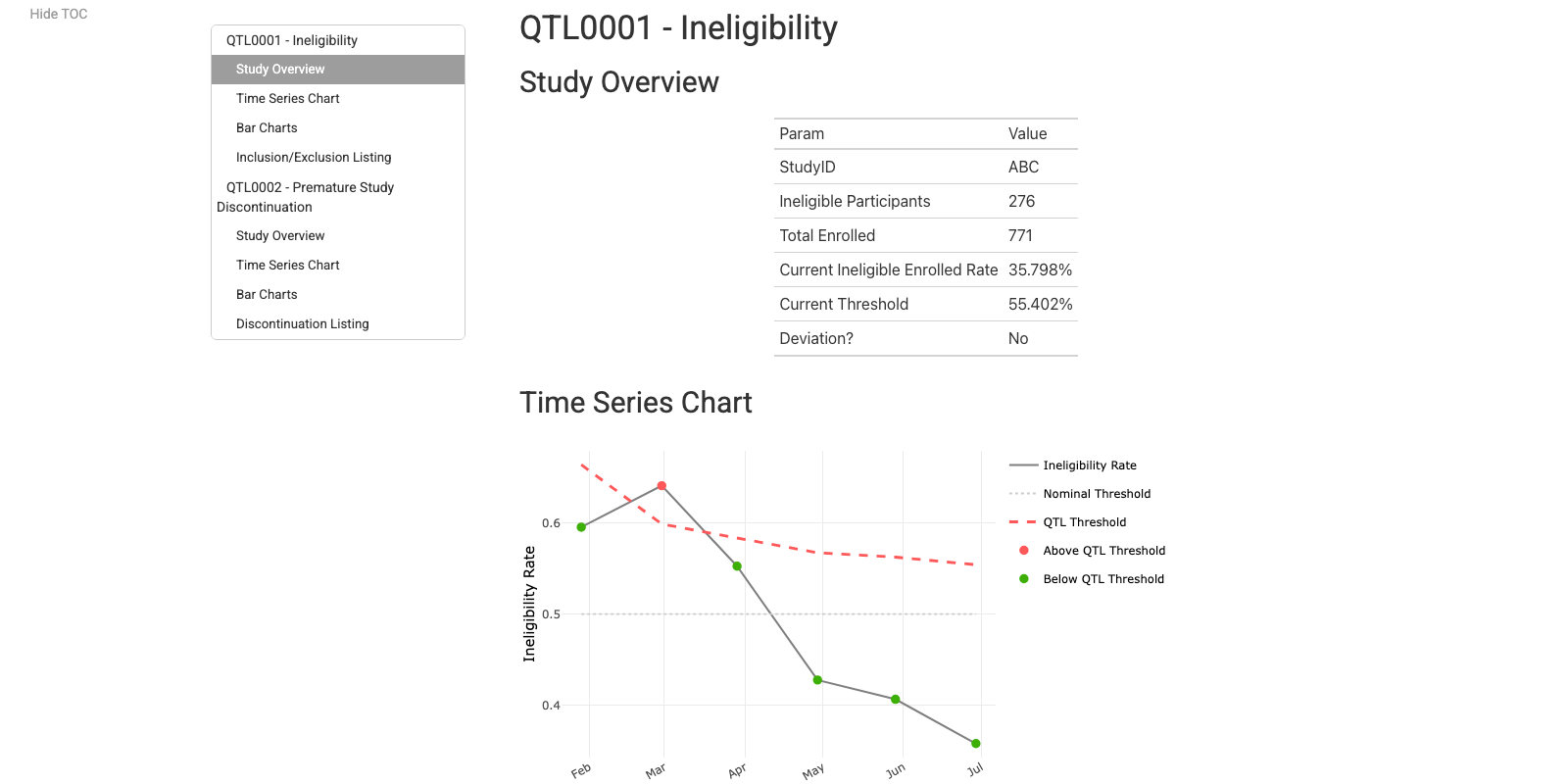
{gsm} Deep Dive App
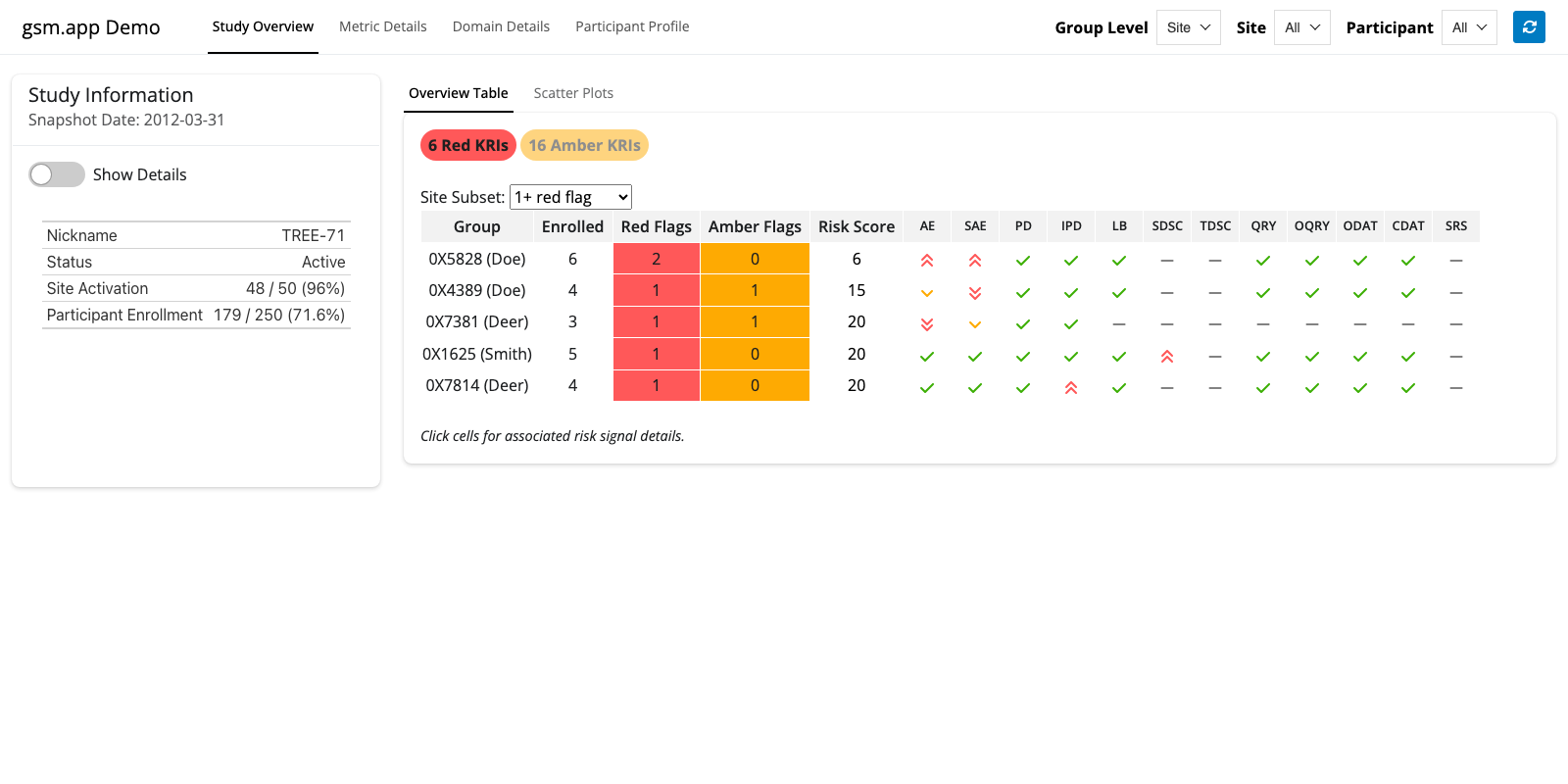
Full {gsm} Data Model
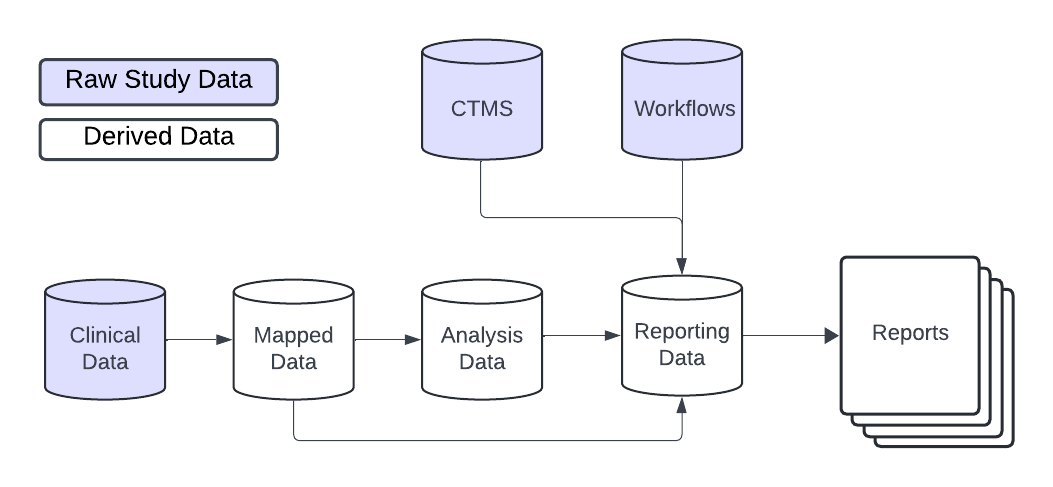
{gsm} ❤️ {workr}
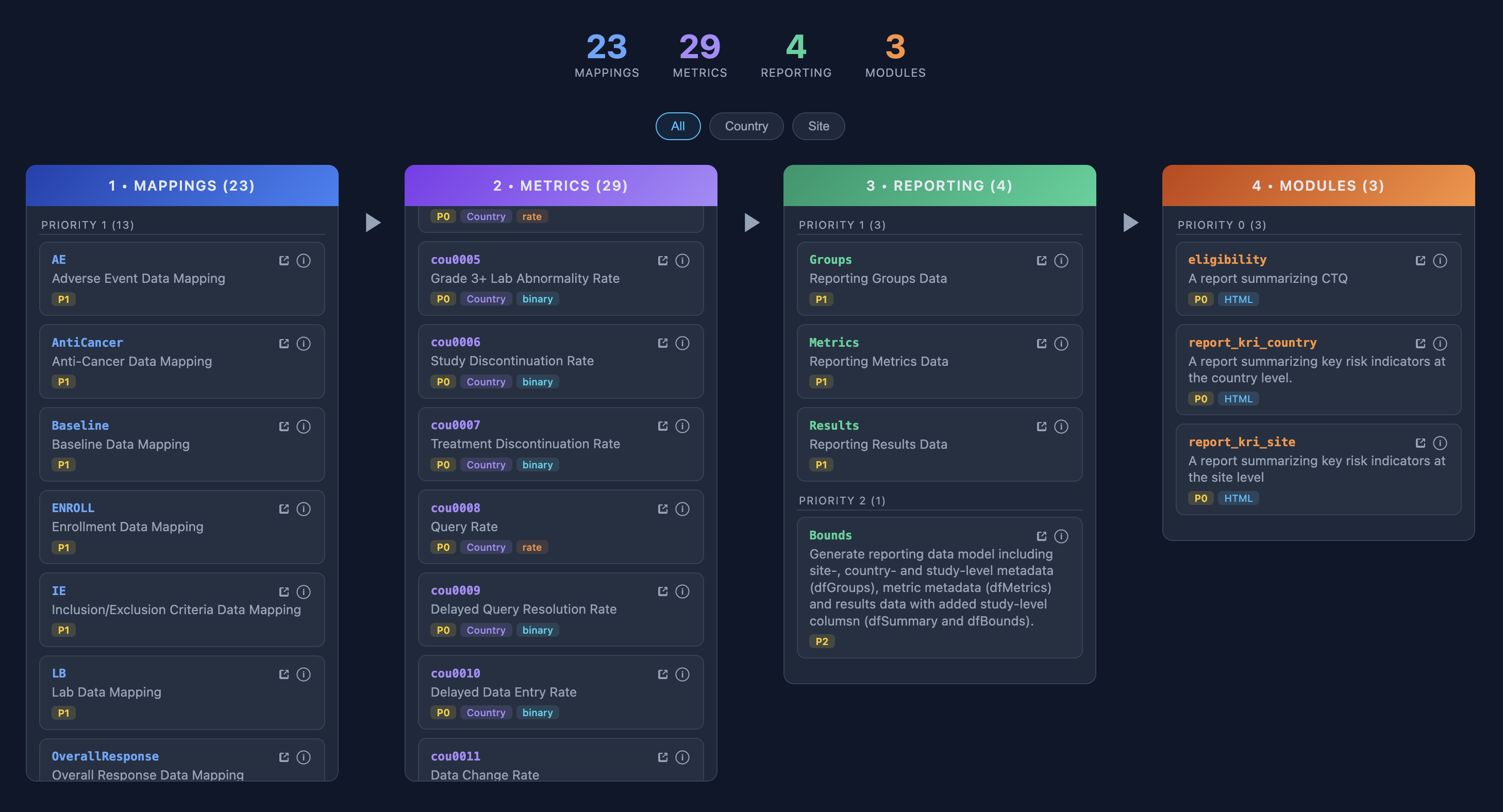
{gsm} + {workr} Study Operations
workr::snapshot()combines workflows across packages- Quarterly snapshots of {gsm} workflows
rvfor managing package versions
- GitHub repo for each study
- Clone default workflows, customize as needed
workr::runProject()runs folders of workflows- Schedule monthly and/or trigger as needed
- Load/save hooks for preferred locations
- folder, database, S3, etc.
GxP Considerations
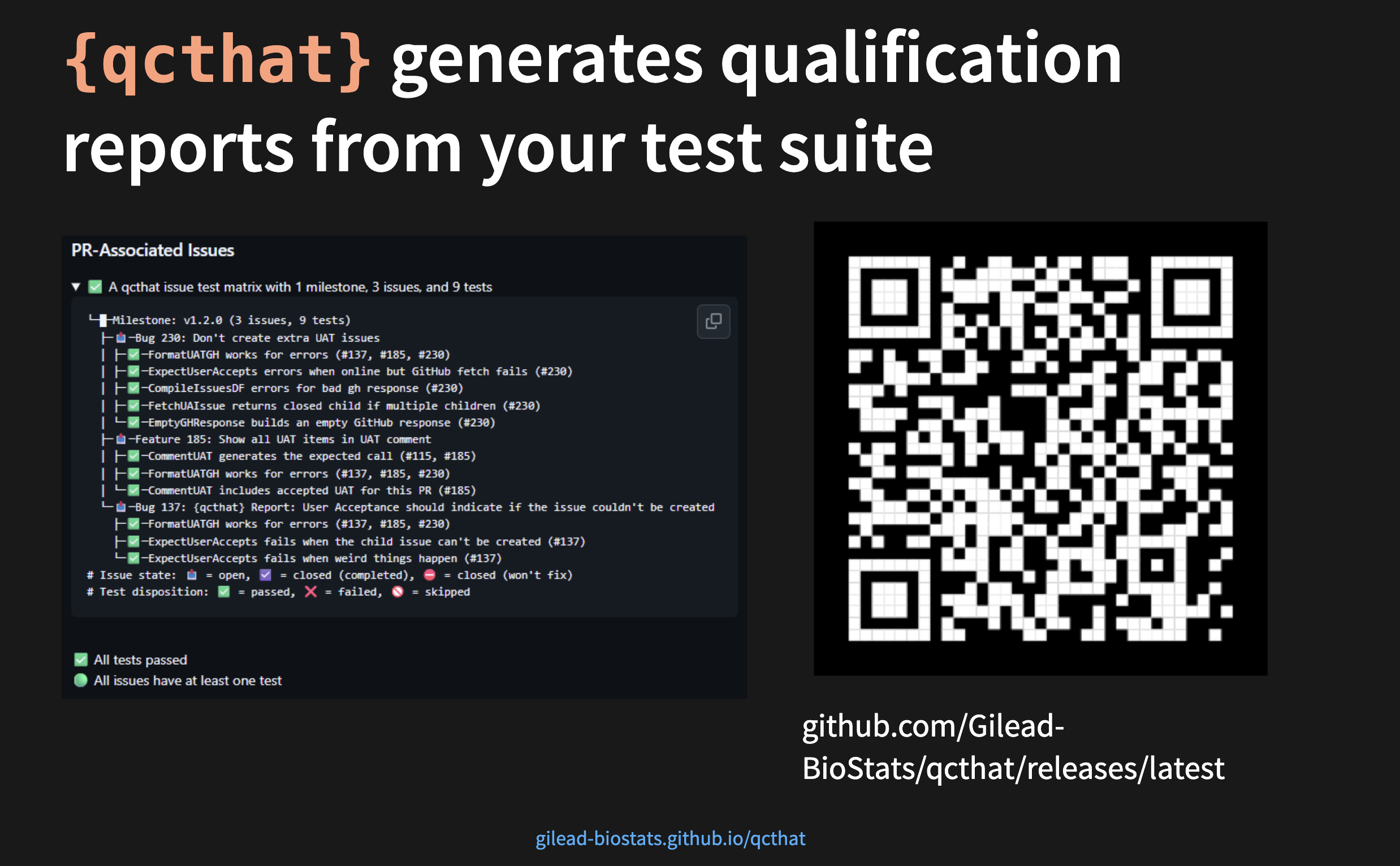
- Extensive qualification via {qcthat}
- {workr}-driven Action Log
- SMEs assess and address detected risks
- In use on 20+ studies.
San Antonio | 1:30-2:30 PM
{workr} ❤️ {pharmaverse}
Not just for ClinOps!
From Raw to SDTM
- {sdtm.oak} is a popular R package that modularizes SDTM programming that is EDC/data-standards agnostic
- The algorithms and sub-algorithms provided can be reused across multiple SDTM domains
- We aim to replicate the results of this vignette
![]() using workflows!
using workflows!
SDTM Example: RAW to VS
What you would run in R
wf <- yaml::read_yaml(system.file("demo_gsmpharmaverse/workflows/1_RAW_TO_SDTM/VS.yaml"))
lData <- list(
dm_raw = read.csv(system.file("raw_data/dm.csv", package = "sdtm.oak")),
vs_raw = read.csv(system.file("raw_data/vitals_raw_data.csv", package = "sdtm.oak")),
study_ct = read.csv(system.file("raw_data/sdtm_ct.csv", package = "sdtm.oak"))
)
RunWorkflow(
lWorkflow = wf,
lData = lData
)What’s happening inside
meta:
ID: VS
Type: SDTM
Description: Transform Raw VS to SDTM VS following sdtm.oak article
Priority: 1
spec:
# Read in data
vs_raw:
_all:
required: true
study_ct:
_all:
required: true
steps:
# Create oak_id_vars
- output: vs_raw2
name: sdtm.oak::generate_oak_id_vars
params:
raw_dat: vs_raw
pat_var: "PATNUM"
raw_src: "vitals"=
SDTM Example: Naming convention considerations
It is important to define how interim objects are handled within lData. As in pipe-based workflows (%>% and |>), teams can either preserve each step by assigning a new output name or overwrite the existing object; in this example, that means choosing vs_raw2 for traceability or overwriting vs_raw.
SDTM Example: CT assignment Steps
Using workflows
# Map topic variable SYSBP and its qualifiers.
- output: vs_sysbp
name: sdtm.oak::hardcode_ct
params:
raw_dat: vs_raw
raw_var: "SYS_BP"
tgt_var: "VSTESTCD"
tgt_val: "SYSBP"
ct_spec: study_ct
ct_clst: "C66741"
- output: vs_sysbp
name: workr::RunQuery
params:
df: vs_sysbp
strQuery: "SELECT * FROM df WHERE VSTESTCD IS NOT NULL"
# Map topic variable SYSBP and its qualifiers.
- output: vs_sysbp
name: sdtm.oak::hardcode_ct
params:
tgt_dat: vs_sysbp
raw_dat: vs_raw
raw_var: "SYS_BP"
tgt_var: "VSTEST"
tgt_val: "Systolic Blood Pressure"
ct_spec: study_ct
ct_clst: "C67153"
- output: vs_sysbp
name: sdtm.oak::assign_no_ct
params:
tgt_dat: vs_sysbp
raw_dat: vs_raw
raw_var: "SYS_BP"
tgt_var: "VSORRES"Using pipes
# Map topic variable SYSBP and its qualifiers.
vs_sysbp <-
hardcode_ct(
raw_dat = vs_raw,
raw_var = "SYS_BP",
tgt_var = "VSTESTCD",
tgt_val = "SYSBP",
ct_spec = study_ct,
ct_clst = "C66741"
) %>%
dplyr::filter(!is.na(.data$VSTESTCD)) %>%
hardcode_ct(
raw_dat = vs_raw,
raw_var = "SYS_BP",
tgt_var = "VSTEST",
tgt_val = "Systolic Blood Pressure",
ct_spec = study_ct,
ct_clst = "C67153",
id_vars = oak_id_vars()
) %>%
assign_no_ct(
raw_dat = vs_raw,
raw_var = "SYS_BP",
tgt_var = "VSORRES",
id_vars = oak_id_vars()
)Each pipe statement is equivalent to a step in the workflow; integration of any package would mostly be depend on familiarity with a particular R Package, not necessarily this workr framework.
From SDTM to ADaM
- {admiral} is a popular R package that modularizes ADaM programming with many extension packages that address specific therapeutic area needs
- Here we’ll highlight just a few specific functions, but the documentation and user guides for {admiral} are some of the best amongst all R packages
ADaM Example: VS to ADVS (add MAP)
What you would run in R
wf <- yaml::read_yaml(system.file("demo_gsmpharmaverse/workflows/2_SDTM_TO_ADAM/ADVS.yaml"))
sdtm <- list(
SDTM_DM = arrow::read_parquet(system.file("demo_gsmpharmaverse/data/SDTM/SDTM_DM.parquet", package = "workr")),
SDTM_VS = arrow::read_parquet(system.file("demo_gsmpharmaverse/data/SDTM/SDTM_VS.parquet", package = "workr"))
)
RunWorkflow(lWorkflow = wf, lData = sdtm)What’s happening inside
meta:
ID: ADVS
Type: ADAM
Description: Create Basic ADVS
Priority: 1
spec:
SDTM_DM:
_all:
required: true
SDTM_VS:
_all:
required: true
steps:
- output: advs
name: admiral::derive_vars_merged
params:
dataset: SDTM_VS
dataset_add: SDTM_DM
new_vars: !expr exprs(TRT01A)
by_vars: !expr exprs(STUDYID, USUBJID)
- output: advs
name: dplyr::mutate
params:
.data: advs
PARAMCD: !expr rlang::expr(.data[["VSTESTCD"]])
AVAL: !expr rlang::expr(.data[["VSORRES"]])
- output: advs
name: admiral::derive_param_map
params:
dataset: advs
by_vars: !expr exprs(STUDYID, USUBJID, TRT01A, VSDTC, VISIT, VISITNUM, VSTPT, VSTPTNUM)
sysbp_code: 'SYSBP'
diabp_code: 'DIABP'
get_unit_expr: !expr rlang::expr(.data[["VSORRESU"]])=
advs <- admiral::derive_vars_merged(
dataset = SDTM_VS,
dataset_add = SDTM_DM,
new_vars = exprs(TRT01A),
by_vars = exprs(STUDYID, USUBJID)
) %>%
mutate(
PARAMCD = VSTESTCD,
AVAL = VSORRES
) %>%
derive_param_map(
by_vars = exprs(STUDYID, USUBJID, TRT01A, VSDTC, VISIT, VISITNUM, VSTPT, VSTPTNUM),
sysbp_code = 'SYSBP',
diabp_code = 'DIABP',
get_unit_expr = VSORRESU
)ADaM Example: Create MAP
- The prior examples establish a consistent workflow pattern for producing an ADaM dataset. In this step,
derive_param_map()derives mean arterial pressure (MAP) fromSYSBPandDIABP. - Compared with stitching together multiple tidyverse transformations, this approach reduces boilerplate and makes the derivation intent clearer.
- A valuable area for future collaboration, particularly in admiral, is harmonizing quasiquotation patterns to improve readability and adoption, especially around
expr(),exprs(), and!!.
From ADaM to TFL
- How every team/organization handles data visualizations may just have the most variance
- We will go over a way that uses a workflow to render Rmarkdown documents with prespecified templates for tables/figures, specifically using
gtsummaryandsafetyCharts - The setup for this would be applicable for shiny apps, static reports, web-based html reports, where to host or view the final object is left to the user
TFL Example: ADVS to TFL
What you would run in R
wf <- yaml::read_yaml(system.file("demo_gsmpharmaverse/workflows/3_ADAM_TO_TFL/WorkProduct1.yaml", package = "workr"), warn = FALSE)
adam <- list(
ADVS = arrow::read_parquet(system.file("demo_gsmpharmaverse/data/ADAM/ADAM_ADVS.parquet", package = "workr"))
)
workr::RunWorkflows(lWorkflows = wf, lData = adam )What’s happening inside
meta:
ID: WorkProduct1
Type: TFL
Description: Create Basic Work Product/Report which can modularize the tables included
Priority: 1
spec:
ADVS:
_all:
required: true
steps:
- output: lParams
name: list
params:
'dfADVS': ADVS
- output: table1
name: rmarkdown::render
params:
input: !expr here::here("demo_gsmpharmaverse", "report_templates", "WorkProduct1.Rmd")
output_file: !expr here::here("demo_gsmpharmaverse", "TFLS", "WorkProduct1.html")
envir: !expr new.env(parent = globalenv())
params: lParams=
lParams <- list(dfADVS = arrow::read_parquet(system.file("demo_gsmpharmaverse/data/ADAM/ADAM_ADVS.parquet", package = "workr")))
table1 <- rmarkdown::render(
input = here::here("demo_gsmpharmaverse", "report_templates", "WorkProduct1.Rmd"),
output_file = here::here("demo_gsmpharmaverse", "TFLS", "WorkProduct1.html"),
envir = new.env(parent = globalenv()),
params = lParams
)TFL Preview
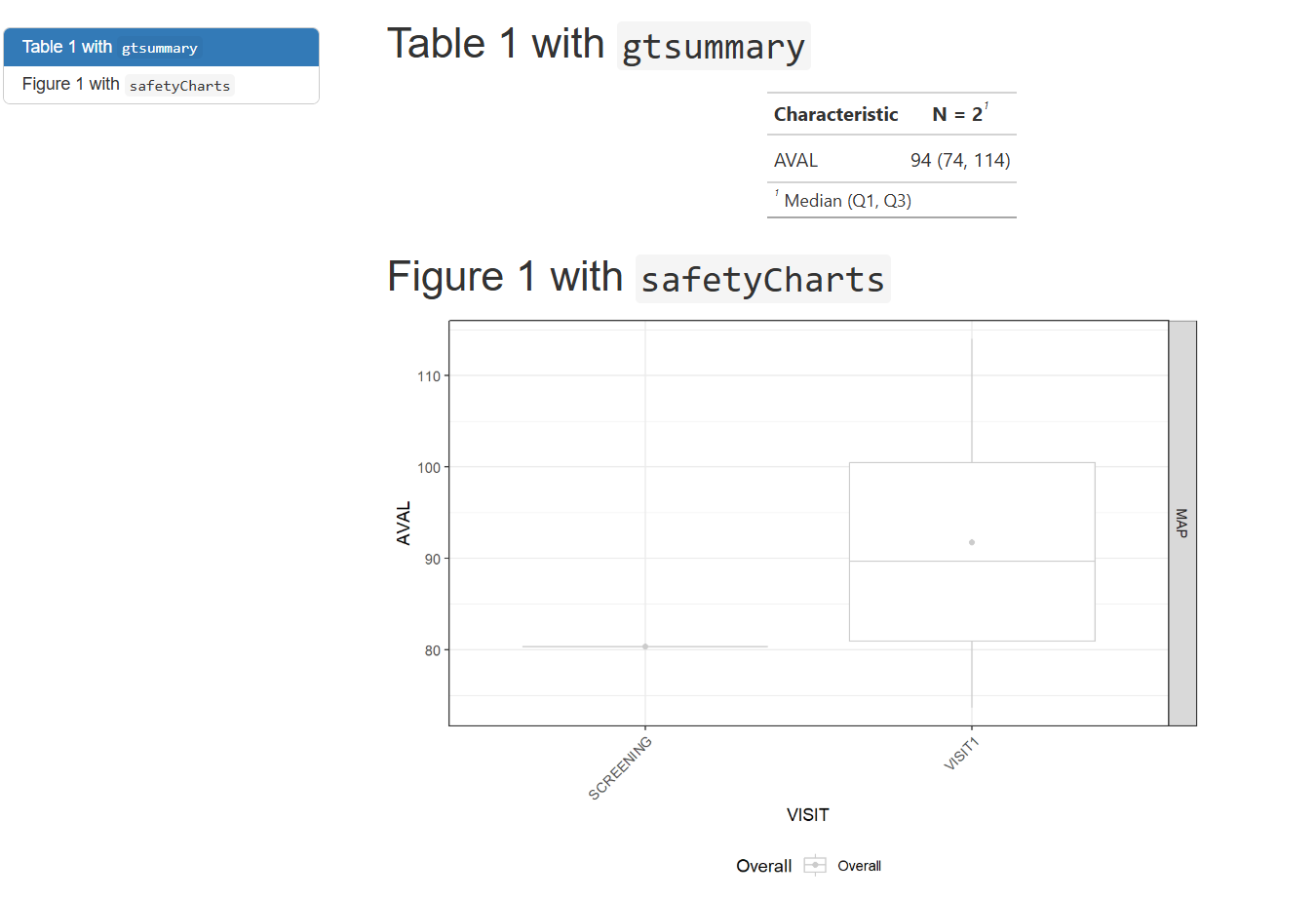
TFL Example: Render Rmarkdown (2)
- These rendered documents (html in this case) can be mounted into a preferred viewing environment (r shiny apps, websites, pdf over email, etc.) for whoever the end user may be.
- The methodology & technical infrastructure will be left to the user.
Parent Rmd 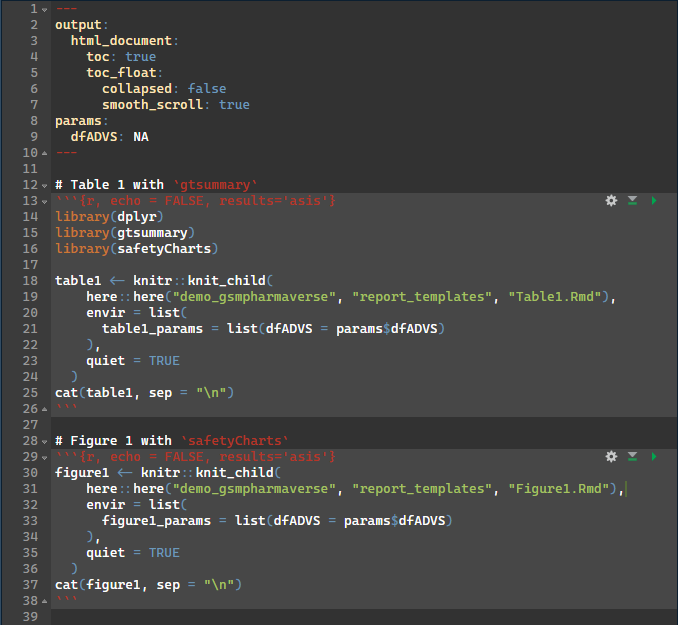
Child Rmd 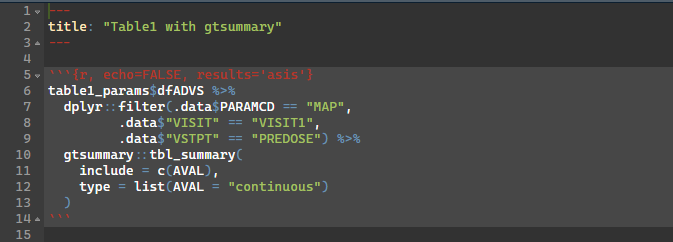
TFL Overview
- Using smaller child .Rmds that may match a respective company standard table/figure template may be favorable in this scenario
- This allows a larger work product/bundle to be stitched together with these child rmarkdowns to reduce clutter of the main document.
- For some deliveries it may be necessary to only have a few tables and figures, and other deliveries a much heftier report. This framework allows a “lego set” style assembled work deliverable in a pick and choose fashion
From ADaM to ARS/ARD
{cards} is an R package for creating CDISC Analysis Results Data (ARD)
It is designed to support automation, reproducibility, reusability, and traceability of analysis results
ARS/ARD Example: ADVS summary
What you would run in R
What’s happening inside
meta:
ID: table_mean_arterial_pressure
Type: ars
Description: Create table 1 ARS
Priority: 1
spec:
ADVS:
_all:
required: true
steps:
- output: predose_visit1_map
name: workr::RunQuery
params:
df: ADVS
strQuery: "SELECT * FROM df WHERE PARAMCD = 'MAP' AND VISIT = 'VISIT1' AND VSTPT = 'PREDOSE'"
- output: table_predose_visit1_map
name: cards::ard_summary
params:
data: predose_visit1_map
variables:
- AVAL=
Final Considerations
The primary outputs (thus far) of these workflows is typically a derived dataset, but persistence (load/save) is intentionally decoupled
Workflows orchestrate transformation logic; storage strategy is flexible and left to the user/organization
Saving outputs (e.g., .csv, .parquet, .json, or a data lake) can be implemented as an additional workflow step/configuration
{workr} Apps
Color demo

Enterprise {workr} Platform gismo
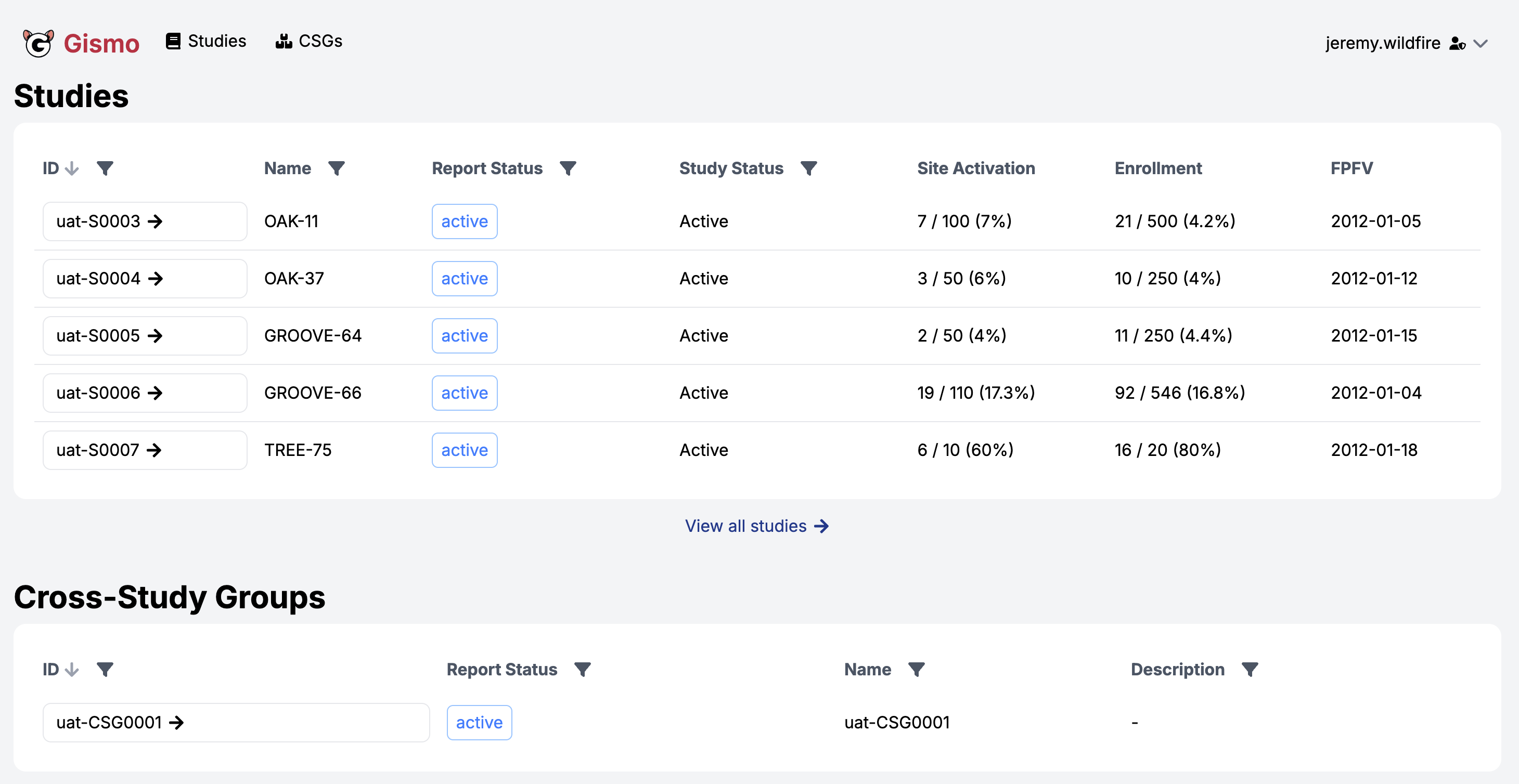
Enterprise {workr} Platform gismo
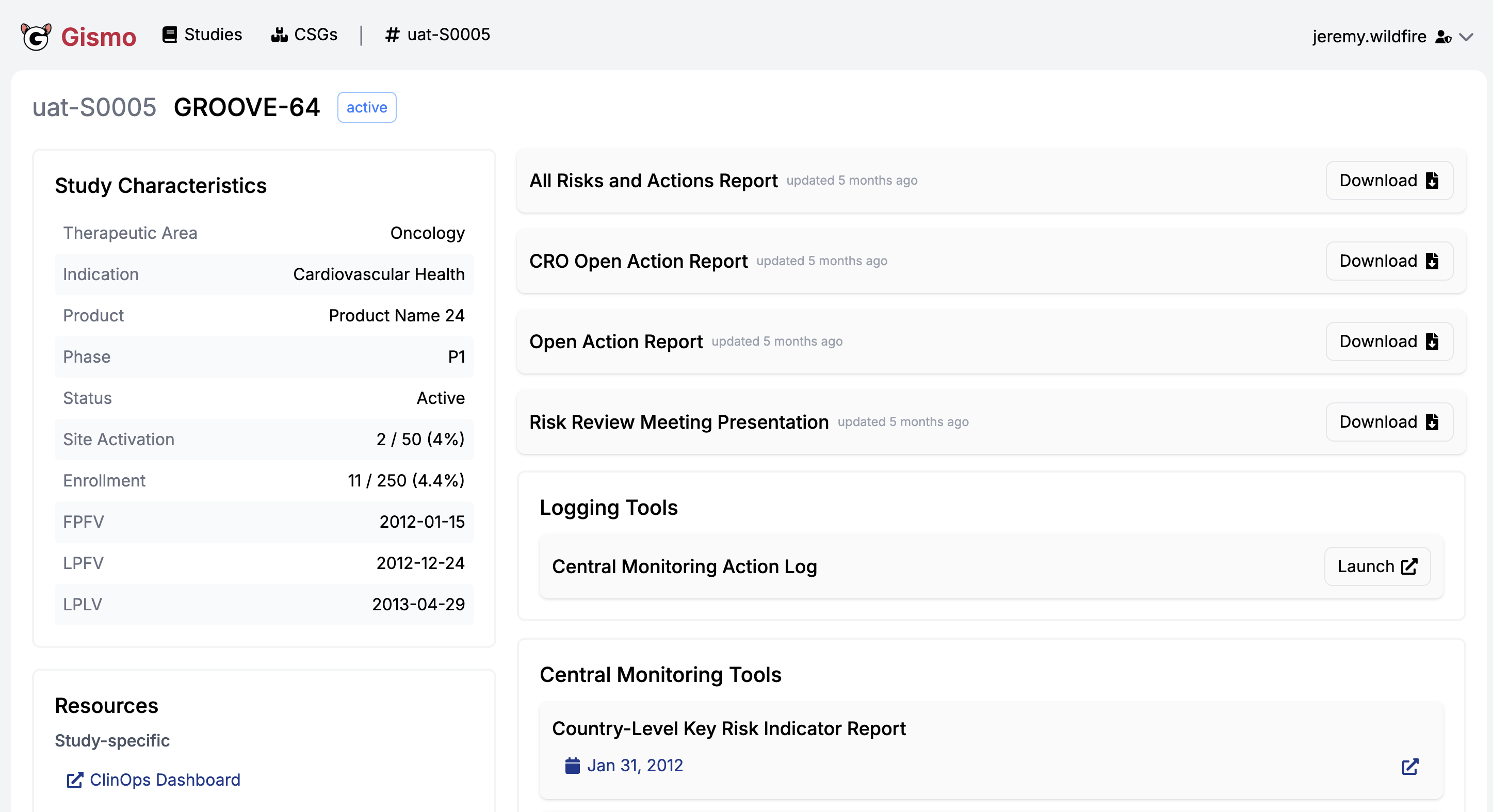
Enterprise {workr} Platform gismo
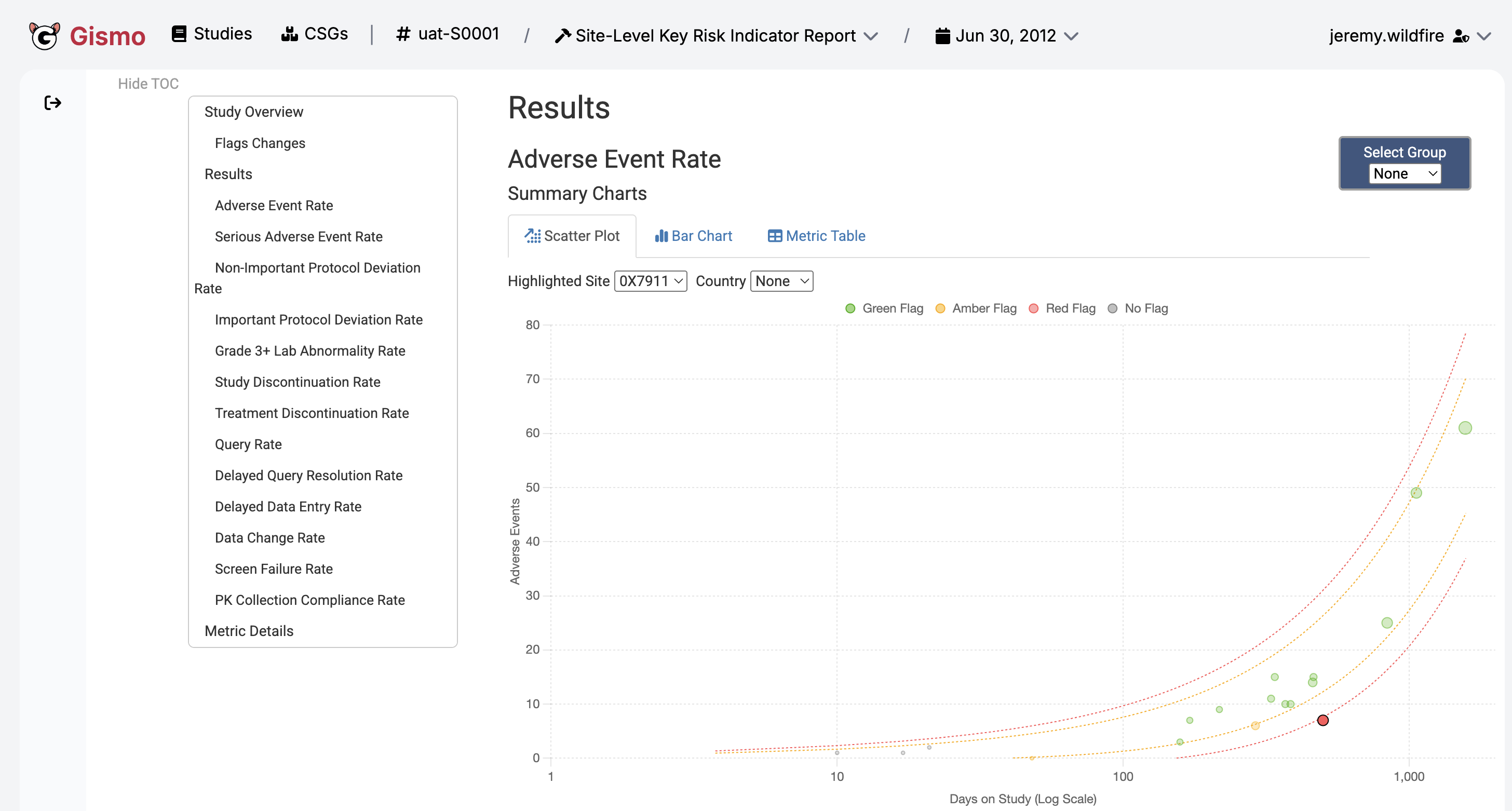
Enterprise {workr} Platform gismo
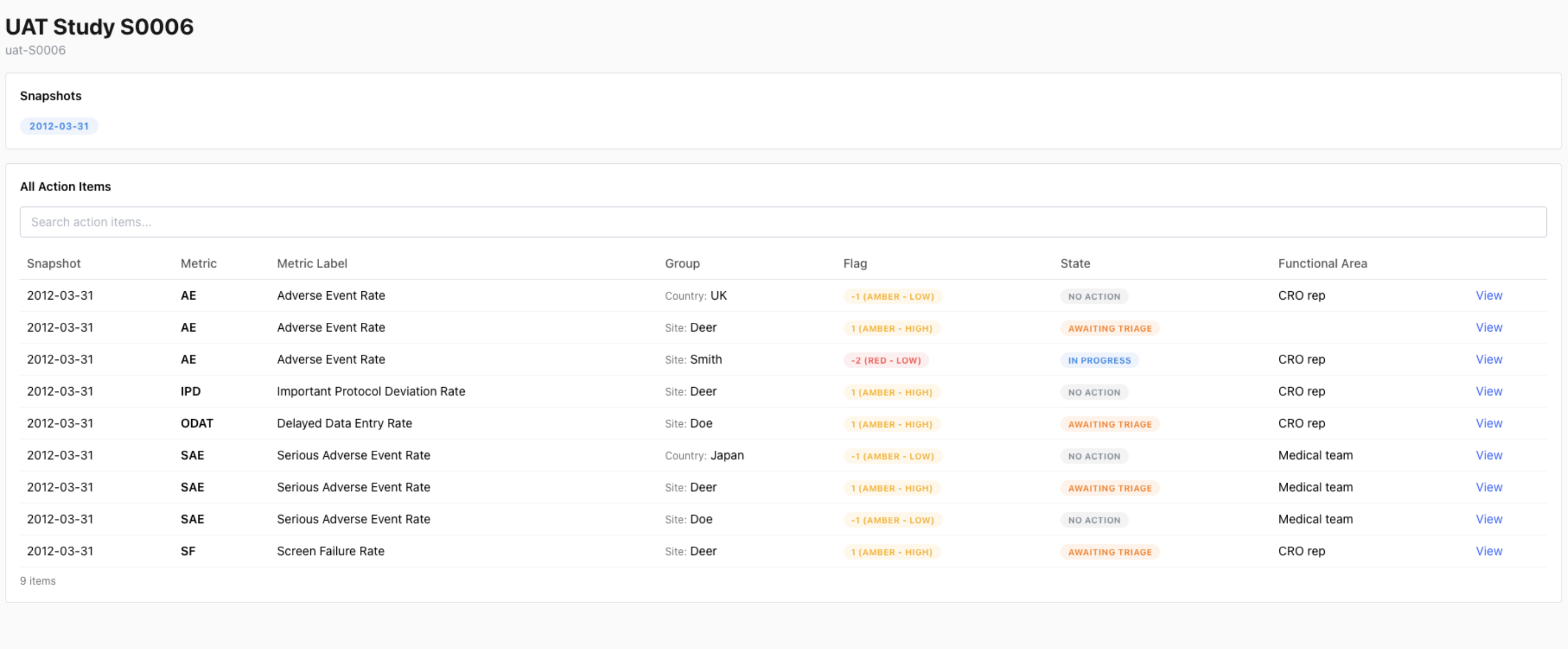
Enterprise {workr} Platform gismo
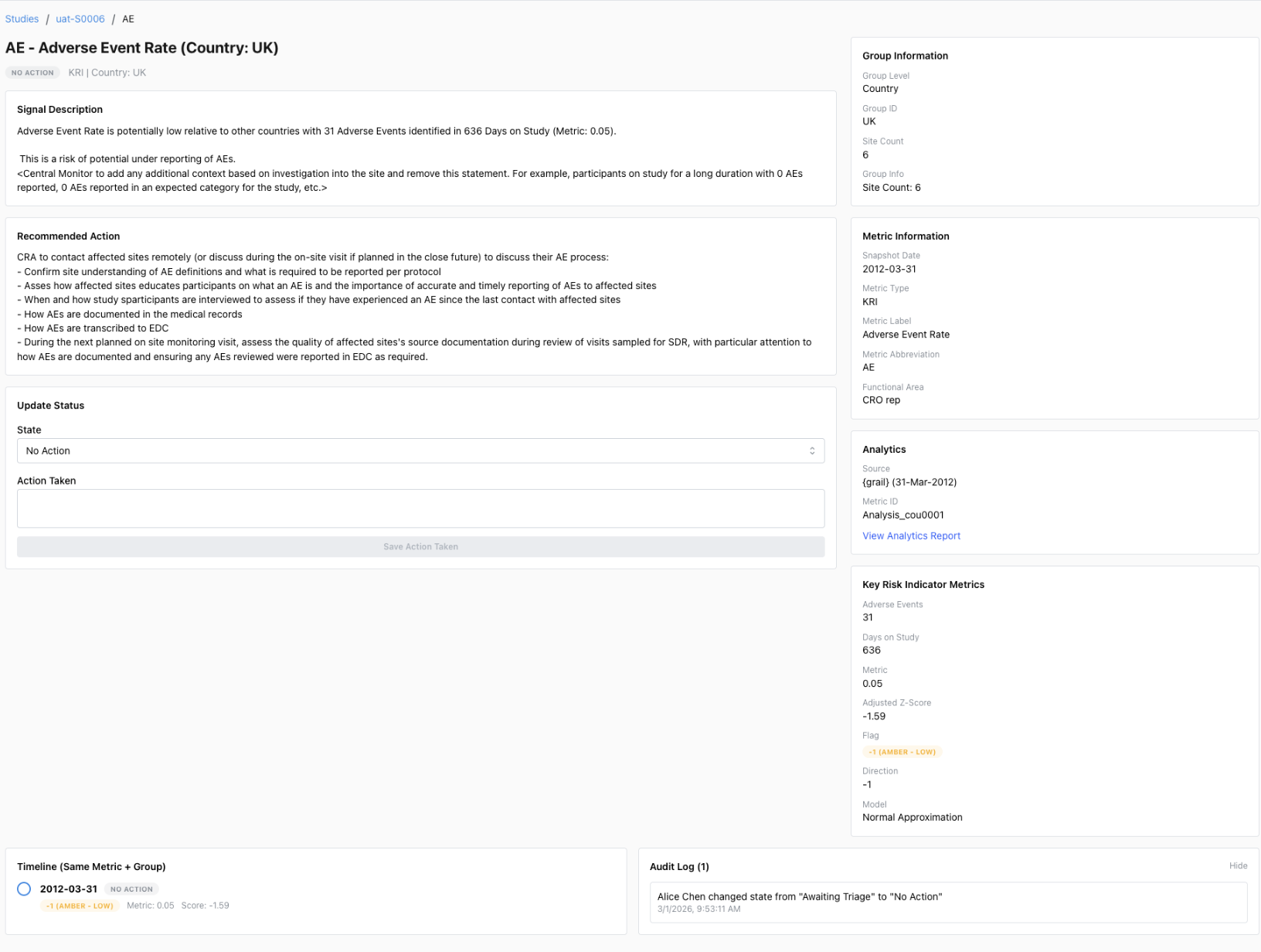
GitHub {workr} Platform open.gismo
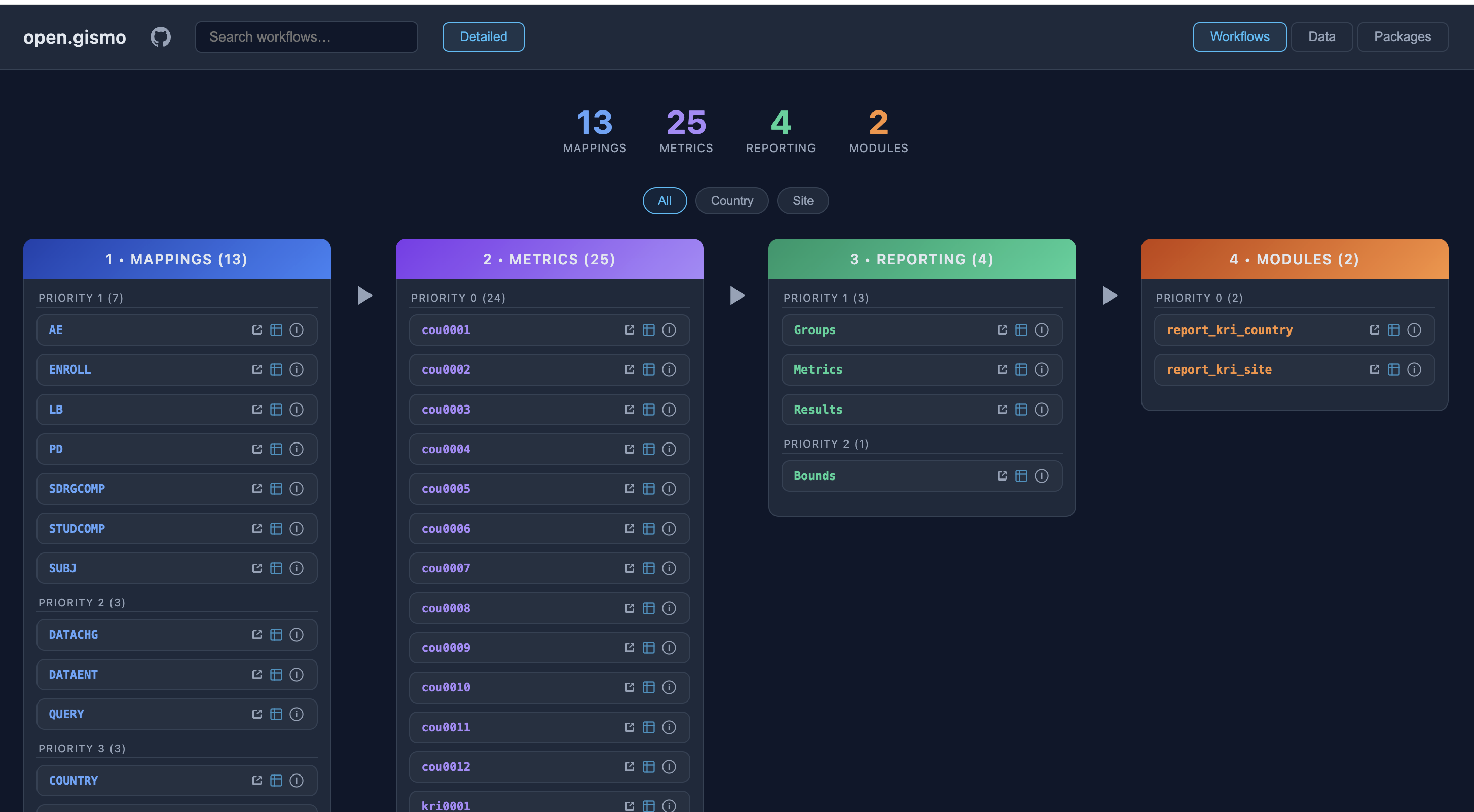
GitHub {workr} Platform open.gismo
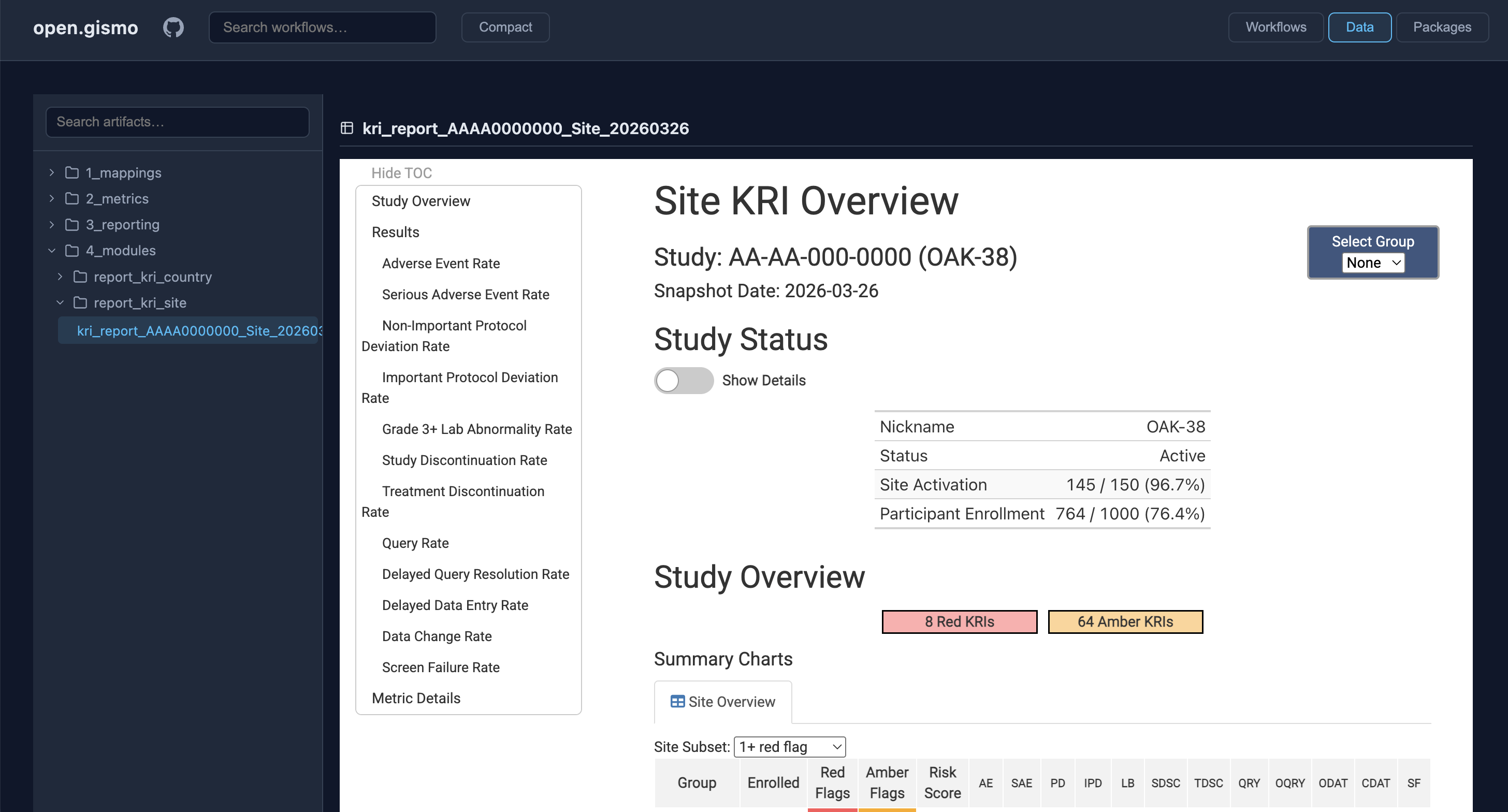
{workr} ❤️ 🤖
- Designed for use with Agentic AI
- Breaks down complex tasks into modular steps
- Agents can generate and execute workflows
- {gsm} and {workr} skills coming soon!
{workr} 🙏 you!
- Simple R workflows
- Highly scalable and flexible
- Designed for GxP use
- Open source and extensible
 using workflows!
using workflows!